Transcribed RNA can be used to study RNA structure and how it relates to function or how proteins and RNA interact. It can also be used for gene silencing using RNAi (studied more often as a possible therapeutic option) or simply serve as a molecular standard in Real-time RT-PCR. Transcribed RNA is also used in Class 2 Clustered Regularly Interspaced Short Palindromic Repeat systems, or CRISPR.
The CRISPR system, which is naturally occurring in bacteria, has been manipulated to perform gene editing in a laboratory environment. To perform CRISPR in the laboratory environment, you need two main reagents:
- The Brains: Guide RNA (gRNA or sgRNA) – Small piece of RNA containing a nucleotide sequence that is capable of binding the chosen Cas Protein, and contains a portion of the sequence that can bind the DNA the researcher intends to modify – the target DNA.
- The Brawn: CRISPR-associated endonuclease (Cas Protein) – The protein that cleaves the target DNA; the most popular Cas protein is called Cas9. The Cas protein is guided by the (gRNA).
Guo et al. used Promega’s RiboMAX™ Large-Scale RNA Production System to produce gRNA to be used in CRISPR for their study to determine the effects of the loss of, or mutations in, a specific gene in fruit flies (1). Atg101 is a gene that plays an important role in autophagy, an intracellular pathway for removing toxins or damaged parts of cells.
Continue reading “Studying Autophagy in Flies Using CRISPR”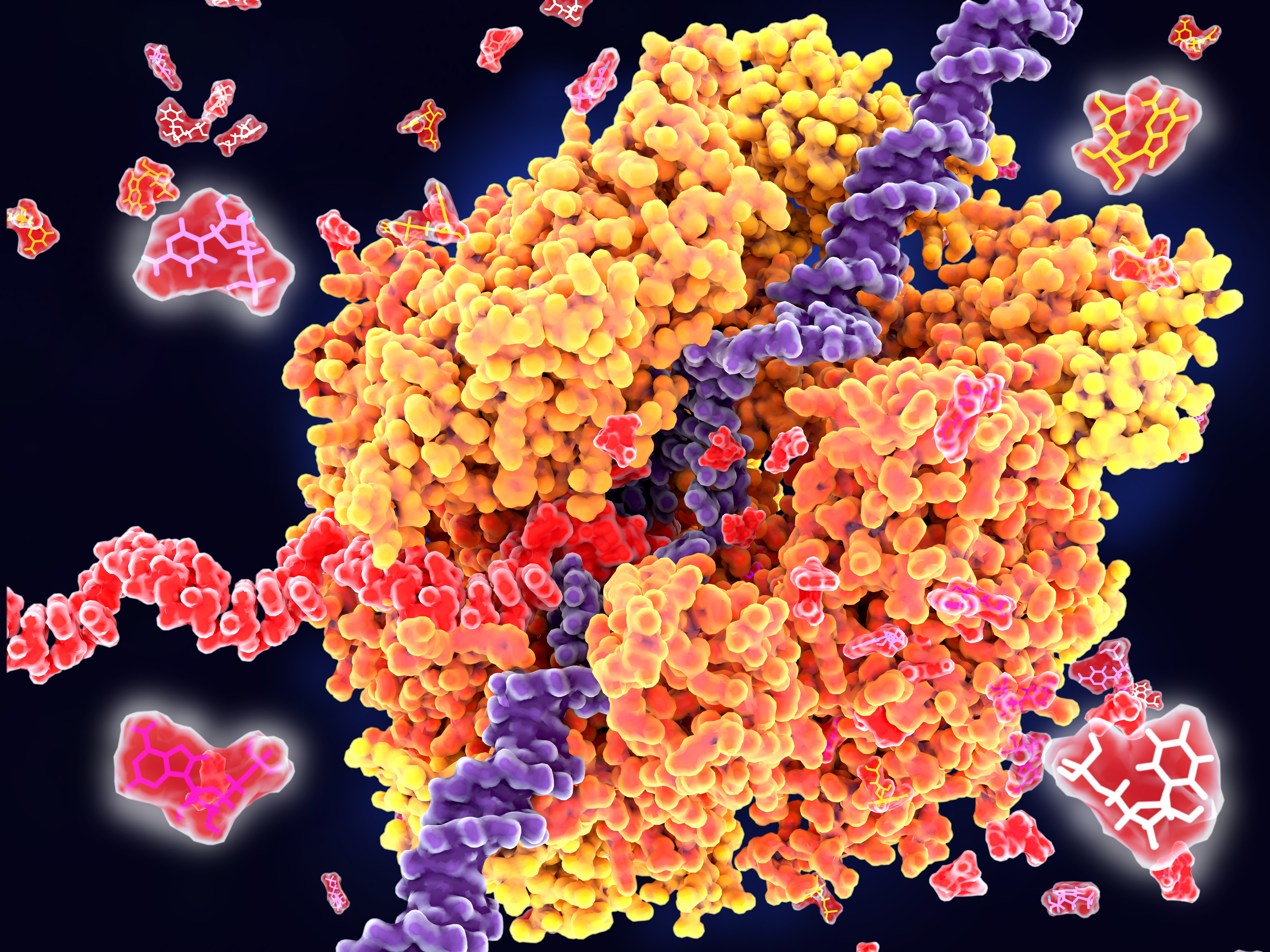

 Scientific inquiry is a process that is revered as much as it is misunderstood. As I listened to
Scientific inquiry is a process that is revered as much as it is misunderstood. As I listened to 

 There are new developments in genetics coming to light every day, each with the potential to dramatically change life as we know it. The increasingly controversial gene editing system, dubbed CRISPR (Clustered Regularly Interspaced Short Palindromic Repeats), is at the root of it all. Harnessed for use in genome editing in 20131, CRISPR has given hope to researchers looking to solve various biological problems. It’s with this technology that researchers anticipate eventually having the means to genetically modify humans and rid society of genetic disorders, such as hemophilia. While this is not yet possible, the building blocks are steadily being developed. Most recently, two groundbreaking studies concerning CRISPR have been released to the public.
There are new developments in genetics coming to light every day, each with the potential to dramatically change life as we know it. The increasingly controversial gene editing system, dubbed CRISPR (Clustered Regularly Interspaced Short Palindromic Repeats), is at the root of it all. Harnessed for use in genome editing in 20131, CRISPR has given hope to researchers looking to solve various biological problems. It’s with this technology that researchers anticipate eventually having the means to genetically modify humans and rid society of genetic disorders, such as hemophilia. While this is not yet possible, the building blocks are steadily being developed. Most recently, two groundbreaking studies concerning CRISPR have been released to the public. 