Why Metabolism Matters in T-Cell Expansion
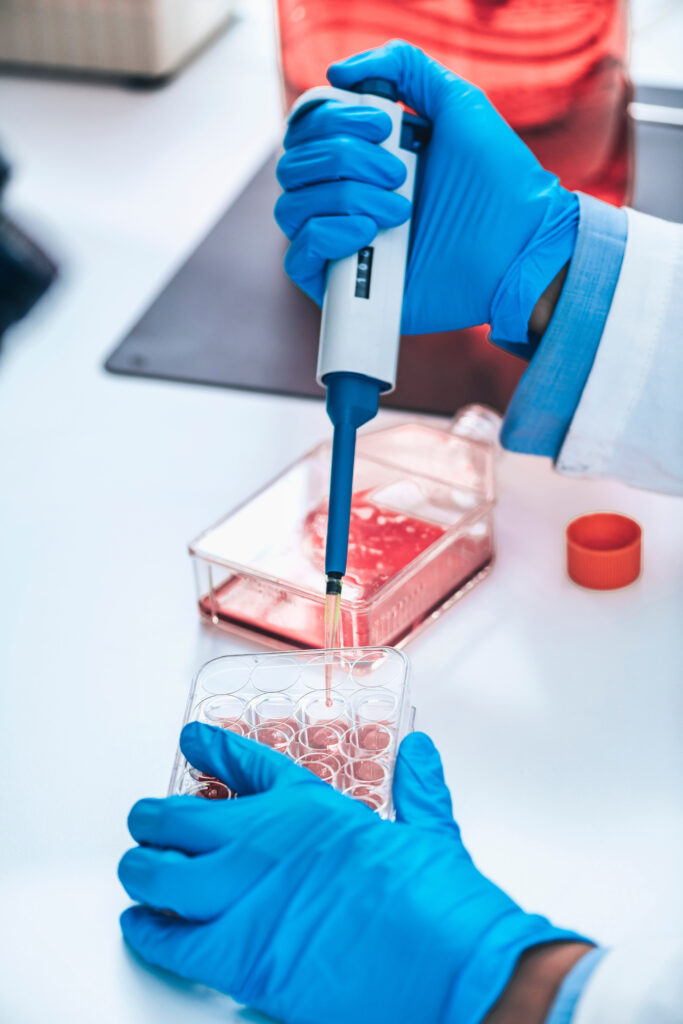
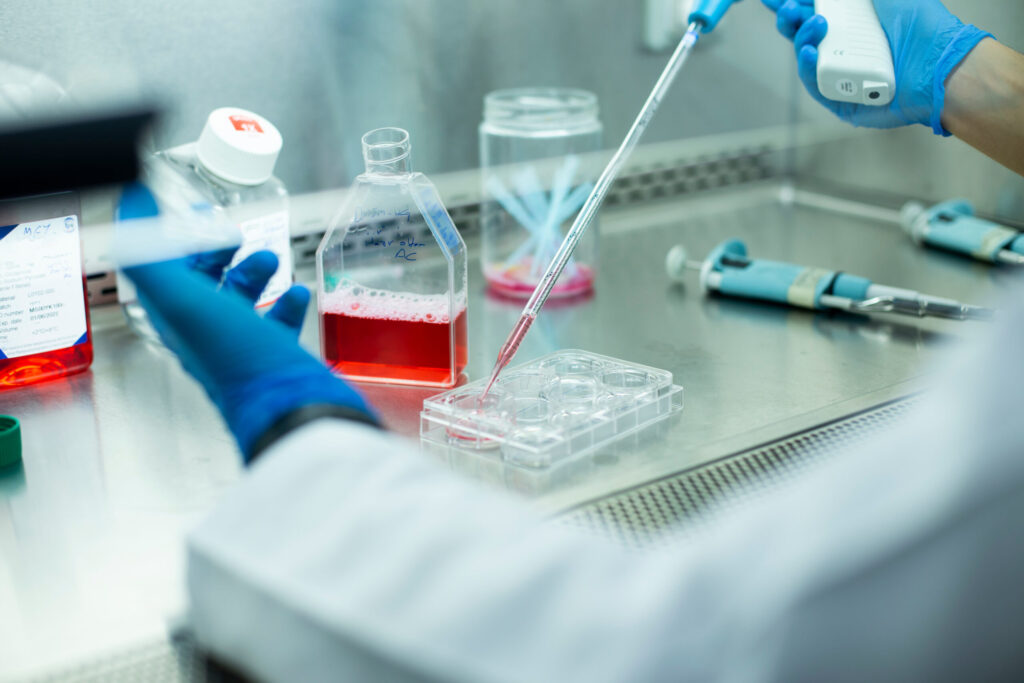
Adoptive T-cell therapies rely on generating metabolically fit, functional cells during ex vivo expansion—but this process often pushes T cells toward highly glycolytic, terminally differentiated states that limit their persistence and therapeutic potential. These metabolic programs begin shifting within hours of activation, therefore understanding early metabolic remodeling is essential for designing culture conditions that support durable, cytotoxic, and memory-enriched T-cell populations.
Researchers at Promega set out to address this challenge by systematically mapping how media composition and activation strength shape T-cell metabolism during the first 72 hours after stimulation. Using a suite of bioluminescent assays, they profiled intracellular energy cofactors, redox balance, and extracellular metabolites across several conditions. This approach revealed distinct, media-driven metabolic states that not only emerged early but also predicted downstream expansion, proliferation, and cytotoxic function.
Their work demonstrates how integrating metabolic profiling into in vitro expansion workflows can provide a more informed framework for optimizing T-cell manufacturing strategies.
Continue reading “Your Media Choice Might Be Designing Your T-Cell Fate”