Limitations of Traditional Protein Study Methods
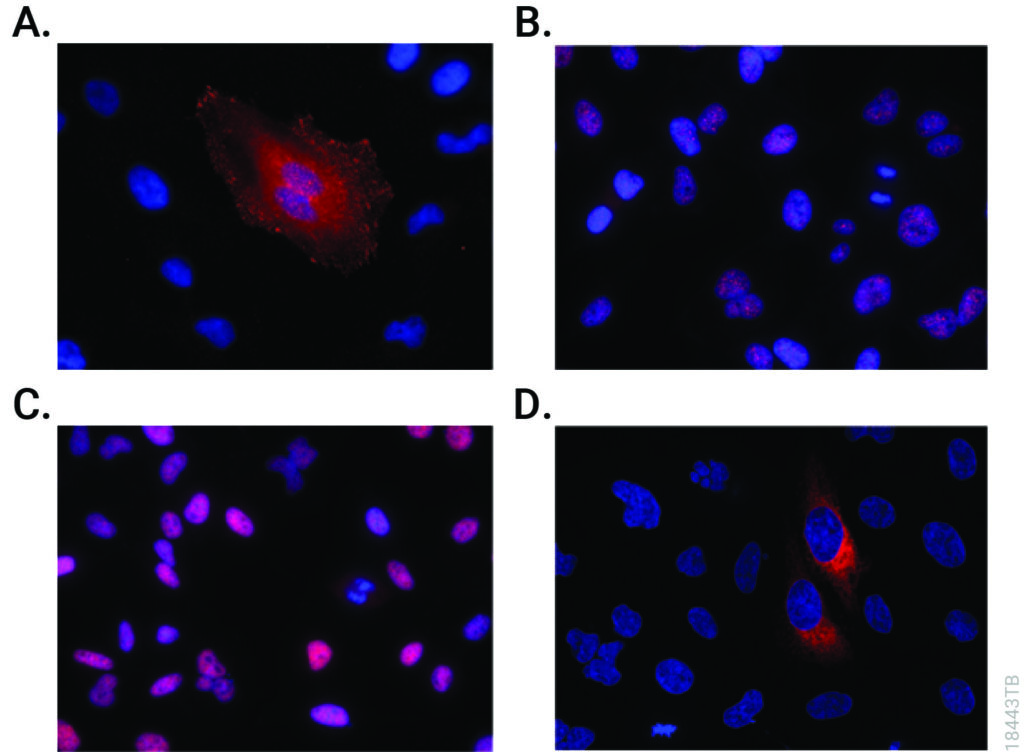
Studying proteins in their native biological context has long been a major challenge in molecular biology. Traditional methods, although widely used, often distort the actual cellular environment and limit functional interpretation. Techniques like antibody-based detection or plasmid-driven overexpression can introduce artifacts and do not allow real-time analysis in living cells.
In this context, the need for tools that enable the observation of proteins as they naturally occur, under physiological conditions, and within live cells is becoming increasingly evident in molecular biology.
Continue reading “CRISPR/Cas9 Endogenous Tagging in Drug Discovery”