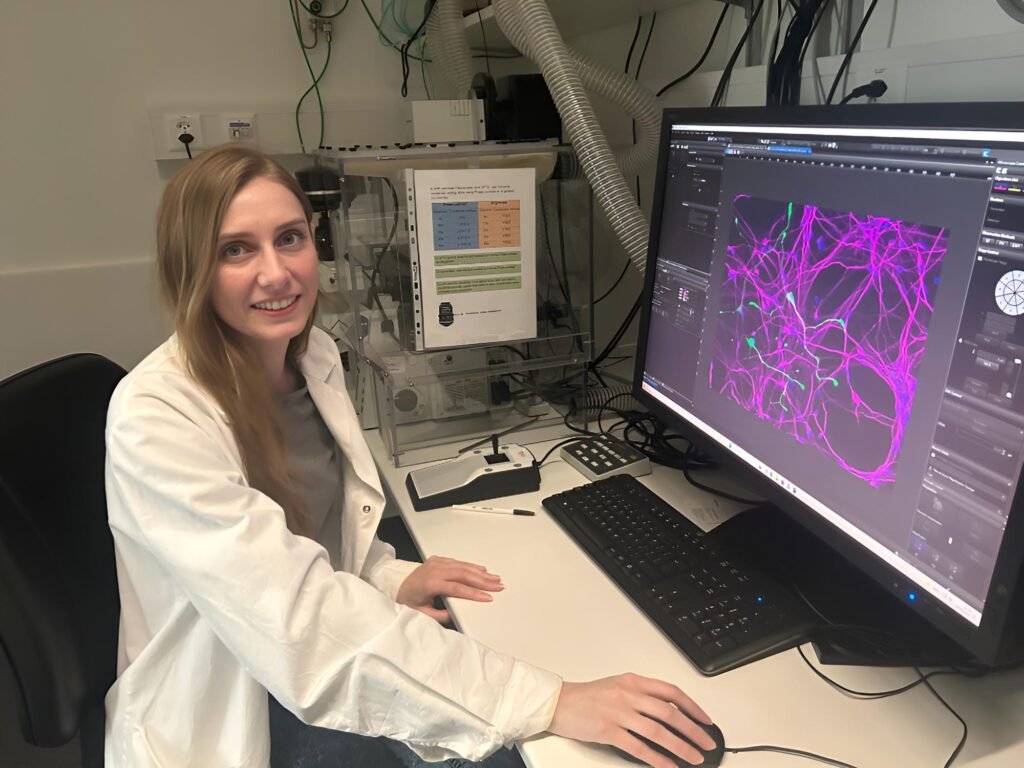
This guest blog post is written by Tian Yang, Associate Product Manager at Promega.
There are often challenges with translating results from a test tube into a living system, demanding more physiologically relevant assays. In drug discovery, demonstrating a compound’s ability to modulate its target protein in live cells is a critical step in the hit-to-lead workflow. A variety of cell-based assays can be used to assess a compound’s activity in live cells. Take kinase inhibitors as an example, these assays can range from substrate phosphorylation assays that more directly report on the activity of target kinases, to genetic reporter assays or cell viability assays that assess the downstream effects of target modulation.
In the case of Tivantinib, several pieces of data from its development were used to establish its role as an inhibitor of MET kinase. MET Kinase is a prominent target for anti-cancer therapeutics due to frequent MET dysregulation in a wide range of tumors. For example, over-activation of MET drives cancer proliferation and metastasis. In the initial report on Tivantinib, in addition to biochemical activity assays performed with purified MET, the activity of Tivantinib in cells was verified by several methods, including: 1) inhibition of phosphorylation of MET and downstream signaling pathways, 2) cytotoxicity in cancer cell lines expressing MET, and 3) antitumor activity in xenograft mouse models (1). Additionally, a co-crystal structure of the MET-Tivantinib complex was solved, seemingly confirming that Tivantinib is a bona fide MET inhibitor capable of engaging MET in live cells (2). Based on these observations and other pre-clinical data, Tivantinib appeared to be a promising drug candidate and was taken through phase 3 clinical trials targeting cancers with MET overexpression. However, Tivantinib ultimately was not approved as a new therapeutic, failing to show efficacy in these phase 3 clinical trials (3,4).
Within three years of the initial publication on Tivantinib, two separate articles challenged the mechanism of action in Tivantinib-induced cytotoxicity of tumor cells (5,6). Authors for both articles showed that Tivantinib can kill both MET-addicted and nonaddicted cells with similar potency. Both articles also concluded that perturbation of microtubule dynamics, instead of MET inhibition, is likely responsible for the cytotoxicity observed with Tivantinib. Considering the failed clinical trials and uncertainties regarding the mechanism of action, one may wonder if the original pre-clinical work adequately determined if Tivantinib effectively binds and inhibits MET in cells? If Tivantinib’s cellular engagement to MET was assessed directly rather than by MET phosphorylation analysis, would a different pre-clinical recommendation have been made?
Continue reading “Drug Target Confirmed? Tivantinib’s Lesson on the Importance of Cellular Target Engagement”