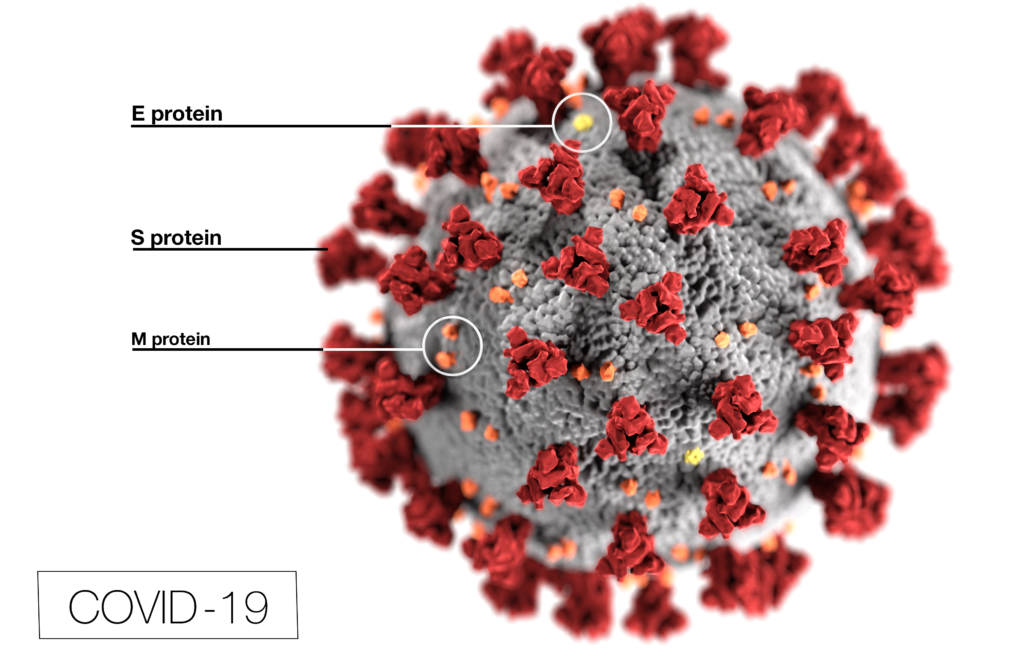
Coronavirus (CoV) researchers are working quickly to understand the entry of SARS-CoV-2 into cells. The Spike or S proteins on the surface of a CoV is trimer. The monomer is composed of an S1 and S2 domain. The division of S1 and S2 happens in the virus producing cell through a furin cleavage site between the two domains. The trimer binds to cell surface proteins. In the case of the SARS-CoV, the receptor is angiotensin converting enzyme 2. (ACE2). The MERS-CoV utilizes the cell-surface dipeptidyl peptidase IV protein. SARS-CoV-2 uses ACE2 as well. Internalized S protein goes though a second cleavage by a host cell protease, near the S1/S2 cleavage site called S2′, which leads to a drastic change in conformation thought to facilitate membrane fusion and entry of the virus into the cell (1).
Rather than work directly with the virus, researchers have chosen to make pseudotyped viral particles. Pseudotyped viral particles contain the envelope proteins of a well-known parent virus (e.g., vesicular stomatitis virus) with the native host cell binding protein (e.g., glycoprotein G) exchanged for the host cell binding protein (S protein) of the virus under investigation. The pseudotyped viral particle typically carries a reporter plasmid, most commonly firefly luciferase (FLuc), with the necessary genetic elements to be packaged in the particle.
To create the pseudotyped viral particle, plasmids or RNA alone are transfected into cells and the pseudotyped viruses work their way through the endoplasmic reticulum and golgi to bud from the cells into the culture medium. The pseudoviruses are used to study the process of viral entry via the exchanged protein from the virus of interest. Entry is monitored through assay of the reporter. The reporter could be a luciferase or a fluorescent protein.
Firefly Luciferase Measurement
Pseudotyped viruses have been used to study host cell entry mediated by the SARS-CoV-2 S protein, as well as S-proteins derived from other coronaviruses. Walls, et al. (1) chose to measure the firefly luciferase (FLuc) reporter with the ONE-Glo™ EX Luciferase Assay System. Letko, Marzi and Munster (2) chose the Bright-Glo™ Luciferase Assay System. Ou, et al. (3) chose the Steady-Glo® Luciferase Assay System and Xia, et al.(4) chose the original Luciferase Assay System. While each firefly luciferase assay performs well, each were designed for different applications.
Xia, et al. chose the original Luciferase Assay System, a first-generation luciferase assay. The assay requires washing of cells and use of either the Cell Culture Lysis Reagent or the Reporter Lysis Buffer to create a lysate. A portion of the lysate is mixed with the Luciferase Assay Reagent, and luminescence measured with a single-tube luminometer, like the GloMax® 20/20 Instrument. The light output from the system is extremely bright but has a half-life of <10 minutes. If you are performing single assays with a single tube luminometer, this is not an issue. Moving to a 96-well plate requires a plate-reading luminometer with injectors. Each individual well must be injected and measured before moving to another well. The process can take up to 1 hour per plate.
The second-generation assays, the “Glo” assays, were designed for multi-well plate applications. These reagents will both lyse the cells and provide the substrates to measure firefly luciferase. The reagent is added to the entire plate and the plate is read quickly one well after the other. On instruments like the GloMax® Discover, an entire 96-well plate is read in about a minute.
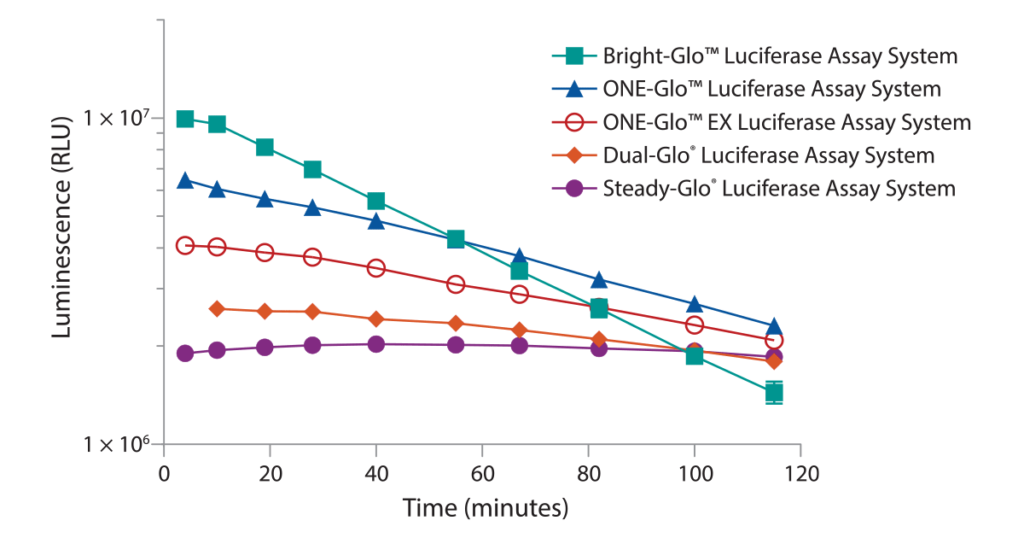
The Steady-Glo® Assay was designed for ultra-high throughput applications where plates are processed in batches. The luminescent signal is quite low in comparison to the 1st generation assays but the signal lasts for hours instead of <10 minutes. The reagent is added to 10s–100s of plates, one after the other, then moved to a luminometer and read, one after the other. The Bright-Glo™ Assay produces a bright luminescent signal much like the 1st generation assay with the added benefit of a ≥25 minute signal half-life. The Bright-Glo™ Assay was designed for continuous process where reagent is added, and the plate is read before moving to the next plate. The ONE-Glo™ and ONE-Glo™ EX Assays give a signal half-life in between Bright-Glo™ and Steady-Glo™ Assays. The ≥50 minute signal half-life allows for continuous or small batch processing of plates. The ONE-Glo™ EX Assay is a more ambient room temperature stable formulation. A comparison of these 2nd generation firefly luciferase assays regarding brightness and signal half-life is shown in Figure 1.
Renilla Luciferase Measurement
Pseudotyped viral particles have also been produced with Renilla luciferase (RLuc) reporters. Pfaender, et al. (6) chose the RLuc reporter for pseudotyped CoV virus production and the Renilla Luciferase Assay System for the luminescent measurement. The Renilla Luciferase Assay is a first-generation assay requiring lysate production prior to addition of the assay reagent. The assay is also low throughput with the same limitations of the Luciferase Assay System for measuring firefly luciferase.
A 2nd generation assay is available called the Renilla-Glo™ Luciferase Assay System. This 2nd generation assay will produce a slightly lower initial signal, but the signal half-life will be ≥60 minutes.
Two reagents are available for kinetic, live-cell measurement of RLuc activity—EnduRen™ and ViviRen™ Live Cell Substrates. Both are modified coelenterazine molecules that become bioactivated within cells to form coelenterazine substrate. Both can be added to the culture media and measured at any time appropriate for the reagent. EnduRen™ Substrate will produce a moderate signal but that signal will be suitable for experiments lasting 24 hours. ViviRen™ Substrate will produce a bright signal but decays more rapidly and should be used for experiments of less than 1 hour.
NanoLuc® Luciferase
NanoLuc® Luciferase is small (19kDa) enzyme molecularly evolved from a deep sea shrimp and, when combined with the optimized substrate furimazine, is ~100x brighter than either FLuc or RLuc(6). The small luciferase is widely cited for the production of recombinant viruses (7) including the MERS (8), SARS-CoV(8) and SARS-CoV-2(9). Work with the NLuc-SARS-CoV and NLuc-MERS-CoV provided early evidence that remdesivir could be a treatment for COVID-19 (10).
Robbiani, et al. (11), at Rockefeller University, chose to create pseudotyped viruses expressing the SARS-CoV-2 S protein using a NLuc reporter in an HIV-based viral particle to gain higher sensitivity for high-throughput applications. The pseudotyped NLuc virus was used to evaluate convalescent serum from over 100 individuals for neutralizing antibodies. The Rockefeller group helped Lui, et al. (12) demonstrate that dimeric ACE2 was necessary for binding the trimeric S protein on SARS-CoV-2. Two ACE2 extracellular domains attached to an IgG Fc region could block the pseudovirus, whereas monomeric ACE2 extracellular domains could not. Interestingly, the cells in the NLuc-expressing pseudovirus studies were lysed with the Cell Culture Lysis Reagent before mixing with the Nano-Glo® Luciferase Assay System.
Interested in tools and resources for viral research? Visit our web page on SARS-CoV-2 Research, Vaccine, and Therapeutic Development.
References
- Walls, A.C., et al. (2020) Structure, function and antigenicity of the SARS-CoV-2 spike glycoprotein. Cell. 181, 281-292.
- Letko, M., Marzi, A. and Munster, V. (2020) Functional assessment of cell entry and receptor usage for SARS-CoV-2 and other lineage B betacoronaviruses. Nat. Microbiol. 5, 562-9.
- Ou, X., et al. (2020) Characterization of spike glycoprotein of SARS-CoV-2 on virus entry and its immune cross-reactivity with SARS-CoV. Nat. Comm. 11, 1620.
- Xia, S., et al. (2020) Inhibition of SARS-CoV-2 infection (previously 2019-nCoV) by a highly potent pan-coronavirus fusion inhibitor targeting its spike protein that harbors a high capacity to mediate membrane fusion. Cell Res. 30, 343-55.
- Pfaender, S., et al. (2020) LY6E impairs coronavirus fusion and confers immune control of viral disease. bioRxiv posted 7 Mar 2020.
- Hall, M.P., et al. (2012) Engineered luciferase reporter from a deep sea shrimp utilizing a novel imidazo-pyrazinone substrate. ACS Chem. Biol. 7, 1848-57.
- Hooper, K. (2018) Size does matter: NanoLuc® Technologies advance virology research.
- Sheahan, T.P., et al. (2017) Broad-spectrum antiviral GS-5734 inhibits both epidemic and zoonotic coronaviruses. Sci. Transl. Med. 9, eaal3653.
- Dinnon 3rd., K.H., et al. (2020) A mouse-adapted SARS-CoV-2 model for the evaluation of COVID-19 medical countermeasures. bioRxiv posted 7 May 2020.
- Hooper, K. (2020) Investigation of remdesivir as a possible treatment for SARS-2-CoV (2019-nCoV). Promega Connections blog.
- Robbiani, D.F., et al. (2020) Convergent antibody responses to SARS-CoV-2 infection in covalescent individuals. bioRxiv posted 22 May 2020.
- Lui, I., et al. (2020) Trimeric SARS-CoV-2 Spike interacts with dimeric ACE2 with limited intra-Spike avidity. bioRxiv posted 21 May 2020.

One thoughtful comment