In a study published in Proceedings of the National Academy of Sciences USA article, Wang et al. used the principle of the Promega NanoBRET™ assay to understand how ERK1/2 phosphorylation of Rabin8, a guanine nucleotide exchange factor, influenced its configuration and subsequent activation of Rab8, a protein that regulates exocytosis.
Rab8 is a member of the Rab family of small GTPases and an important regulator of membrane trafficking from the trans Golgi network and recycling endosomes to the plasma membrane. Wang et al. were interested in learning how the guanine nucleotide exchange factor (GEF) Rabin8, a known activator of Rab8, was itself activated to better understand how Rab8 and exocytosis were regulated in the cell. First, they confirmed if the consensus extracellular-signal-regulated kinases ERK1/2 phosphorylation motif uncovered in Rabin8 resulted in phosphorylation of Rabin8. Both in vitro analysis and cell-based assays confirmed that ERK1/2 phosphorylated Rabin8. Next, the GEF activity of Rabin8 was assessed to determine if ERK1/2 phosphorylation activated the GEF. Researchers confirmed activation of Rabin8 GEF in vitro.
How did the phosphorylation of Rabin8 by ERK1/2 activate the GEF catalytic domain? Previous work by Wang et al. using a Rabin8 mutant suggested that Rabin8 might adopt an autoinhibitory configuration by having the C terminus block the GEF domain activity. To investigate this hypothesis, a Rabin8 construct was made with NanoLuc® luciferase on the N terminus and HaloTag® protein on the C terminus. A control t-SRARE protein, syntaxin-4 (STX4), known to have a closed conformation, was constructed with the same NanoLuc® luciferase-STX4-HaloTag® protein configuration. The NanoBRET™ assay works when the two proteins, NanoLuc® and HaloTag® are brought into close proximity. The donor NanoLuc® luciferase in the presence of its substrate will emit light. The HaloTag® protein bound to the acceptor HaloTag® NanoBRET™ 618 Ligand if close to the light-emitting NanoLuc® protein will fluoresce. This is the basis of the NanoBRET™ assay. Both the control STX4 protein and Rabin8 were expressed in E. coli and the NanoBRET™ Nano-Glo® Substrate added. For both proteins, fluorescence was detected, and as a negative control, used TEV protease to cleave HaloTag® from the protein fusions and eliminated the NanoBRET™ signal. These results suggested that the N and C terminus of Rabin8 were close together and if the C-terminal HaloTag® protein was removed, this energy transference no longer occurred.
This hypothesis was further explored by comparing the NanoBRET™ signal from Rabin8 with a gain-of-function Rabin8 mutant. The NanoBRET™ assay showed a greater NanoBRET™ ratio for Rabin8 compared to its mutant, reinforcing that the GEF activity is normally blocked by a conformational change in Rabin8. What happens to the NanoBRET™ ratio when Rabin8 is exposed to constitutively active ERK2 or a kinase dead ERK2? Rabin8 with the kinase dead ERK2 had a significantly higher NanoBRET™ ratio than the Rabin8 with active ERK2, indicating phosphorylation played a role in Rabin8’s conformational change. This differential phosphorylation was confirmed with Western blotting. Comparing wildtype Rabin8 with Rabin8-4D, where the serines that are phosphorylated by ERK1/2 were replaced with aspartate, rendering these amino acids phosphorylation mimics, in the NanoBRET™ assay showed a lower NanoBRET™ ratio for Rabin-4D compared to Rabin8. These experiments all suggest that Rabin8 is configured to block the GEF activity until phosphorylated by ERK1/2 when the catalytic domain becomes active. Researchers subsequently show that phosphorylation of Rabin8 promotes its binding to Rab8 and plays a role in recycling to the plasma membrane.
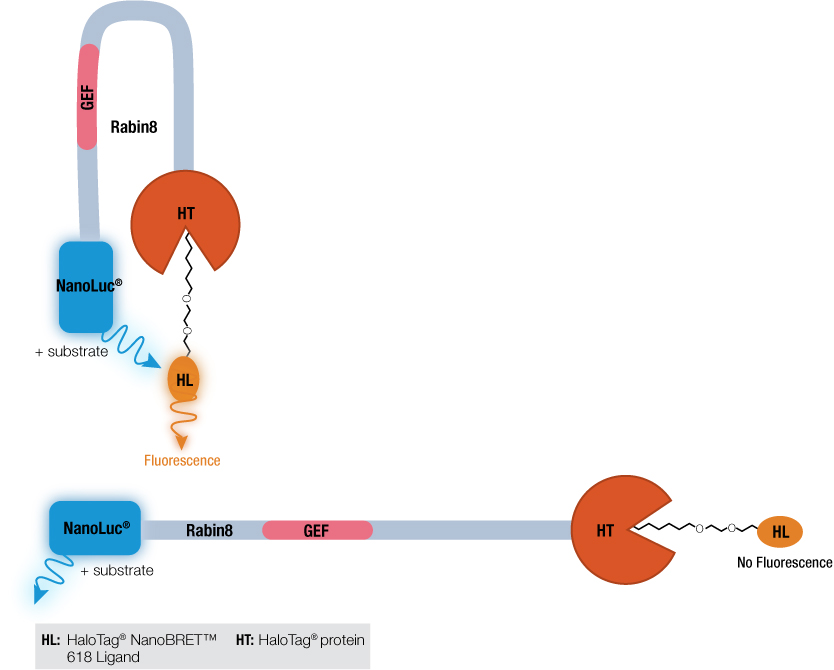
Wang et al. used the NanoBRET™ assay not to look at protein:protein interactions, the typical application of bioluminescence resonance energy transfer (BRET), but to test if a single protein was inhibiting its own catalytic activity. By using attaching the luminescent protein NanoLuc® luciferase to one terminus and the HaloTag® protein to the other, researchers demonstrated that Rabin8 did undergo autoinhibition.
Reference
Wang, J., Wu, B., Feng, S., Cai, G., Tuluc, F., Peränen, J. and Guo, W. (2014) Activation of Rab8 guanine nucleotide exchange factor Rabin8 by ERK1/2 in response to EGF signaling. Proceedings of the National Academy of Sciences 112 (1), 148–53. DOI: http://dx.doi.org/10.1073/pnas.1412089112
Sara Klink
Latest posts by Sara Klink (see all)
- A One-Two Punch to Knock Out HIV - September 28, 2021
- Toxicity Studies in Organoid Models: Developing an Alternative to Animal Testing - June 10, 2021
- Herd Immunity: What the Flock Are You Talking About? - May 10, 2021
