Today’s post was written by guest blogger Anupama Gopalakrishnan, Global Product Manager for the Genetic Identity group at Promega.
Next-generation sequencing (NGS), or massively parallel sequencing (MPS), is a powerful tool for genomic research. This high-throughput technology is fast and accessible—you can acquire a robust data set from a single run. While NGS systems are widely used in evolutionary biology and genetics, there is a window of opportunity for adoption of this technology in the forensic sciences.
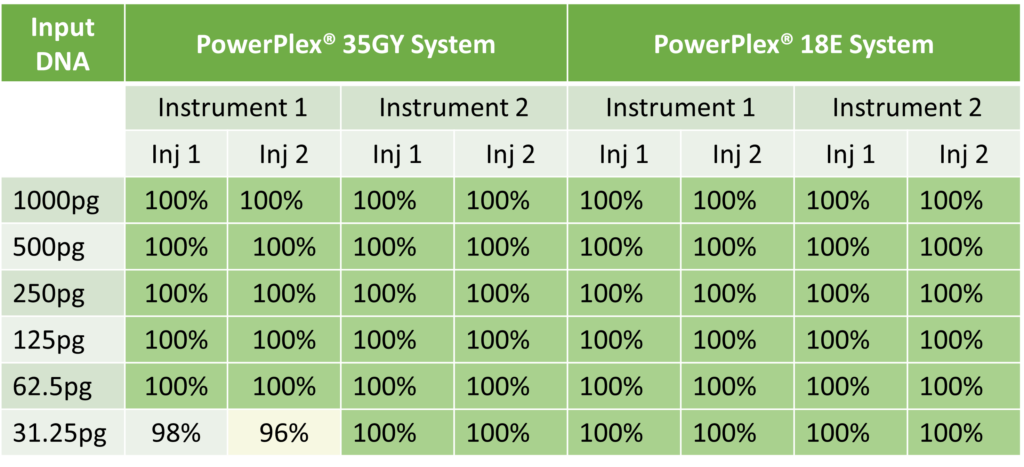
Currently, the gold standard is capillary electrophoresis (CE)-based technologies to analyze short tandem repeats (STR). These systems continue to evolve with increasing sensitivity, robustness and inhibitor tolerance by the introduction of probabilistic genotyping in data analysis—all with a combined goal of extracting maximum identity information from low quantity challenging samples. However, obtaining profiles from these samples and the interpretation of mixture samples continue to pose challenges.
MPS systems enable simultaneous analysis of forensically relevant genetic markers to improve efficiency, capacity and resolution—with the ability to generate results on nearly 10-fold more genetic loci than the current technology. What samples would truly benefit from MPS? Mixture samples, undoubtedly. The benefit of MPS is also exemplified in cases where the samples are highly degraded or the only samples available are teeth, bones and hairs without a follicle. By adding a sequencing component to the allele length component of CE technology, MPS resolves the current greatest challenges in forensic DNA analysis—namely identifying allele sharing between contributors and PCR artifacts, such as stutter. Additionally, single nucleotide polymorphisms in flanking sequence of the repeat sequence can identify additional alleles contributing to discrimination power. For example, sequencing of Y chromosome loci can help distinguish between mixed male samples from the same paternal lineage and therefore, provide valuable information in decoding mixtures that contain more than one male contributor. Also, since MPS technology is not limited by real-estate, all primers in a MPS system can target small loci maximizing the probability of obtaining a usable profile from degraded DNA typical of challenging samples.
Continue reading “Is MPS right for your forensics lab?”