When I was a post-doc at UT Southwestern, I was fortunate to interact with two Nobel prize winners, Johann Deisenhofer and Fred Gilman. Johann once helped me move a -80°C freezer into his lab when we lost power in my building. I once replaced my boss at small faculty mixer with a guest speaker and had a drink with Fred Gilman and several other faculty members from around the university. Among the faculty, one professor had a cell phone on his belt, an odd sight in 1995. Fred Gilman asked him what it was and why he had it. It was so his lab could notify him of good results anytime of the day. Fred laughed and told him to get rid of it – if it’s good data, it will survive until morning.
I was reminded of this story when I read a recent paper by Bodle, C.R. et al (1) about the development of a NanoBiT® Complementation Assay (2) to measure interactions of Regulators of G Protein Signaling (RGS) with Gα proteins in cells. (Fred Gilman was the first to isolate a G protein and that led to him being a co-recipient of the Nobel Prize in 1994). The authors created over a dozen NanoBiT Gα:RGS domain pairs and found they could classify different RGS proteins by the speed of the interaction in a cellular context. The interactions were readily reversible with known inhibitors and were suitable for high-throughput screening due to Z’ factors exceeding 0.5. The study paves the way for future work to identify broad spectrum RGS domain:Gα inhibitors and even RGS domain-specific inhibitors. This is the second paper applying NanoBiT Technology to GPCR studies (3).
A Little Background…
A primary function of GPCRs is transmission of extracellular signals across the plasma membrane via coupling with intracellular heterotrimeric G proteins. Upon receptor stimulation, the Gα subunit dissociates from the βγ subunit, initiating the cascade of downstream second messenger pathways that alter transcription (4). The Gα subunits are actually slow GTPases that propagate signals when GTP is bound but shutdown and reassociate with the βγ subunit when GTP is cleaved to GDP. This activation process is known as the GTPase cycle. G proteins are extremely slow GTPases.
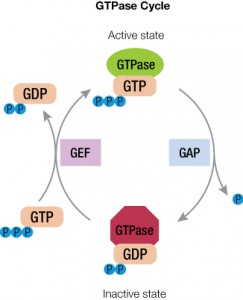
Cells require rapid responses for signal transduction and thus need a way to switch rapidly from the on state to the off state. Accessory proteins assist with this cycle to control how long the G protein is in the active form. These GTPase activating or accelerating proteins (or GAPs) interact with the GTP-bound form and stabilize the transition state to cleave the γ-phosphate of GTP. Another factor, guanine nucleotide exchange factor or GEF, helps remove GDP from the active site to allow GTP to bind again (5).
RGS proteins contain the RGS domain, which serves the role of a GAP from Gαi and Gαq proteins. RGS domains and Gα proteins only interact when GTP is bound accelerating the conversion to GDP which breaks the interaction. Fortunately, an artificial means of forming this interaction is known through the use of a compound called AlF4, which mimics the position of the γ-phosphate of GTP in a GDP-bound Gαi protein and allows interaction with the RGS proteins.
The paper…
Past work on the RGS domain:Gα interactions relied on in vitro methods which brought challenges when moving to a cellular context (e.g., permeability of the compound of interest). Others have applied cell-based assays to study the interactions through secondary downstream events like Ca2+ release. Such cellular studies led to questions as to whether the compound affected the interaction or the downstream assay. Compounds have been discovered that disrupt RGS:Gα interactions using such methods and, coincidentally, work through covalent attachment to cysteine residues on the RGS domain. The authors were looking for a broadly applicable method to assess RGS domain:Gα interactions directly in a cellular context. They turned to the NanoBiT live-cell protein:protein interaction assay (2).
The NanoBiT Assay is a protein complementation assay composed of a large part (17.6kDa or LgBiT) and a small part (11aa or SmBiT), which are fused to two potentially interacting proteins. If the proteins interact, LgBiT and SmBiT are brought together to form active luciferase. LgBiT and SmBiT have low affinity for each other (190µM) and so the binding partners are truly needed to bring them together.
In the Bodle paper, work began by determining the optimal orientation of LgBiT and SmBiT on the two interacting proteins. Four versions of Gαi1 were created (with either LgBiT or SmBiT fused to the N- or C-terminus) and co-transfected with the complementary four versions of the RGS domains derived from either RGS4 or RGS17. The authors chose the best interacting pair based on a combination of signal and fold increase over background in the presence of AlF4. Both studies yielded the identical set-up for maximum signal with fusions at the N-terminus, LgBiT-Gαi1 with SmBiT-RGS4 or RGS17. This gave the authors confidence to build SmBiT-RGS domains with other members of the RGS family (RGS2, 5, 6, 7, 8, 10, 14, 16 and 18) to produce a panel of assays. They also built a version of the RGS4 domain with four cysteines converted to alanine (Cys-less RGS4). RGS2 was an anomoly in this set as RGS2 is known to interact with Gαq, not Gαi1, so the LgBiT-Gαq construct was produced and worked well with SmBiT-RGS2 with no measurable interaction with LgBiT-Gαi1.
The authors followed the kinetics of the interactions and were able to identify two classes of AlF4-induced interaction formation rate. They speculate that the interaction rates could reflect regulation of acute signaling events and sustained signaling events. Interestingly, the Cys-less RGS4 interaction was quite similar to the wild-type RGS4 and the authors commented that such mutants could be useful in identifying non-cysteine reactive inhibitors.
Having demonstrated that the NanoBiT Assay could measure the interactions, the authors investigated whether the interaction could be disrupted. They chose the known RGS4 inhibitor, 6383479, for this test. The kinetic interaction of LgBiT-Gαi1:SmBiT-RGS4 was formed with AlF4 and monitored. At maximal interaction (~40 min), 6383479 was added. The interaction was disrupted and leveled off in about 10 minutes, achieving about 80% signal reduction.
There are eight subfamilies of RGS proteins based on homology in the RGS and accessory domains. One group, including RGS6 and RGS7, contain a G-protein gamma-like (GGL) domain that can interact with the atypical Gβ5 protein and thus participate in other cellular events. A NanoBiT assay was developed to test this interaction of the RGS6 GGL domain with Gβ5. As expected, substitution of the RGS6 RGS domain construct into this interaction assay did not form a complex with Gβ5.
Now having proved that they have a specific interaction assay that can be disrupted, the authors wanted to evaluate if the NanoBiT Assay was robust enough for screening applications by measuring the Z´ factor. Assays were set up in 96-well plates with a weak signal assay, RGS6:Gαi1 for a positive signal and RGS6:NanoBiT Negative Control for a negative signal, and a strong signal assay similarly with RGS8. The Gβ5:RGS6 GGL assay was also measured in a similar way. The weak signal assay reached a Z´ value of 0.5 in 30 minutes following AlF4 addition and maintained >0.5 values for an additional 56 min, plateauing at 0.69. The strong signal assay attained 0.5 in 24 minutes and maintained that >0.5 for an additional 52 min. The Gβ5 assay passed 0.5 in 20 minutes and maintained >0.5 for an additional 102 min, leveled at 0.73. Interestingly, the authors discovered that the RGS:Gα pairs worked best at 2X recommended substrate concentration to generate acceptable Z´ numbers. The RGS GGL:Gβ5 pair worked fine with 1X substrate.
Thus, these authors showed that NanoBiT Technology can be used to create broadly applicable RGS:Gα assays in a cellular context. The formation of the complex is easily reversed by appropriate inhibitors and stands up to the requirements of a high-throughput assay. I look forward to seeing future studies from this lab applying the assays.
References
- Bodle, C.R., et al. (2017) Development of a bimolecular luminescence complementation assay for RGS:G protein interaction in cells. Analytical Biochem., 522, 10-17.
- Dixon, A.S., et al. (2016) NanoLuc complementation reporter optimized for accurate measurment of protein interactions in cells. ACS Chem. Biol., 11, 400-8.
- Canneart, A., et al. (2016) Detection and activity profiling of synthetic cannabinoids and their metabolites with a newly developed bioassay. Analytical Chem., 88, 11476-85. This paper is covered in a previous PromegaConnections blog.
- Cheng, Z., et al. (2010) Luciferase reporter assay system for deciphering GPCR pathways. Curr. Chem. Genomics, 4, 84-91.
- Mondal, S., Hsaio, K. and Goueli, S.A. (2015) A homogeneous bioluminescent system for measuring GTPase, GTPase activating protein and guanine nucleotide exhange factor activities. Assay Drug Dev. Technol., 13, 444-55.
Learn more about NanoBiT Technology.
Post updated April 11, 2024 to address broken links.
