 I should preface this blog by stating that I am a nucleic acids gal. My years in the lab were spent with tubes of DNA and RNA. In fact my one and only tentative foray into the field of protein resulted in a Western Blot so ugly that those who witnessed it have been sworn to secrecy. Given all of this, the mapping of the human proteome might seem like an odd topic for me to write about. Except that it isn’t really, because the sequencing of the proteome offers answers to some of the questions that the sequencing of the genome didn’t.
I should preface this blog by stating that I am a nucleic acids gal. My years in the lab were spent with tubes of DNA and RNA. In fact my one and only tentative foray into the field of protein resulted in a Western Blot so ugly that those who witnessed it have been sworn to secrecy. Given all of this, the mapping of the human proteome might seem like an odd topic for me to write about. Except that it isn’t really, because the sequencing of the proteome offers answers to some of the questions that the sequencing of the genome didn’t.
First, let’s start with what a proteome is: A proteome is all the proteins expressed at a certain time point. It can be as limited as the proteome of a single cell or as all encompassing as the proteome of an entire genome. However, unlike the genome, which is genetic information encoded in an organism’s DNA or RNA, the makeup of a proteome can vary dramatically as a result of expression patterns, alternative splicing events and post-translational modifications.
The genome is a constant, what you see today is what will still be there tomorrow. The proteome, on the other hand, is a constantly changing landscape. Up regulation or down regulation of a gene can mean more or less protein is present. Alternative splicing and post-translational modifications can result in fundamental changes to the protein itself.
In other words, if the genome is a beautiful, pristine Ansel Adams print, then the proteome is that same scene as interpreted by Andy Warhol—in Technicolor and 3D. Continue reading “Discovering the Complexity of the Human Proteome”
Like this:
Like Loading...






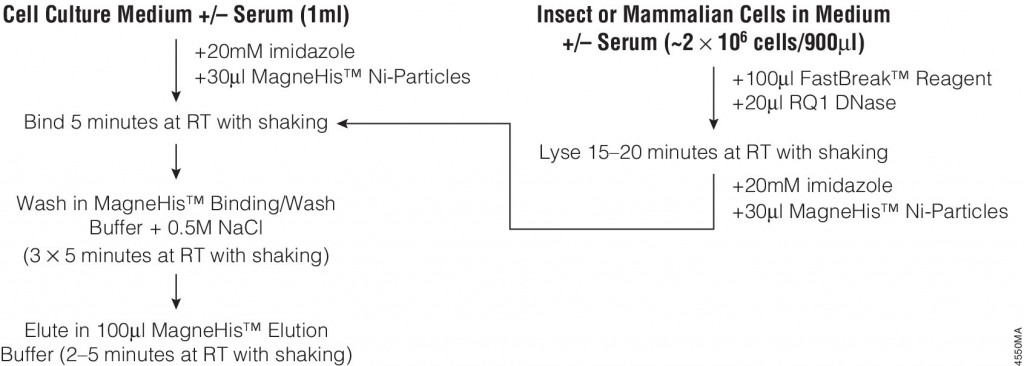
 Many different polypeptide fusion partners or affinity tags have been developed to facilitate purification of target proteins. The most commonly used tag for the purification and detection of recombinant expressed proteins is the His tag. Cloning vectors designed to generate His-tagged proteins contain 5–10 histidine residues at either the C- or N terminus of the expressed protein. The His tag adds only 0.84kDa to the mass of the protein and is nonimmunogenic. Also, because the tertiary structure of the tag is not important for purification, His-tagged proteins can be purified using native or denaturing conditions. The affinity of histidine residues for immobilized nickel allows selective purification of His-tagged proteins. The
Many different polypeptide fusion partners or affinity tags have been developed to facilitate purification of target proteins. The most commonly used tag for the purification and detection of recombinant expressed proteins is the His tag. Cloning vectors designed to generate His-tagged proteins contain 5–10 histidine residues at either the C- or N terminus of the expressed protein. The His tag adds only 0.84kDa to the mass of the protein and is nonimmunogenic. Also, because the tertiary structure of the tag is not important for purification, His-tagged proteins can be purified using native or denaturing conditions. The affinity of histidine residues for immobilized nickel allows selective purification of His-tagged proteins. The