The authors used ultrashort pulsed (USP) lasers in their research as this treatment is known to inactivate a spectrum of bacteria and viruses including nonenveloped viruses, a class of virus that resists inactivation. Furthermore, the laser treatment is nonionizing and does not modify proteins covalently, meaning that proteins present in blood are likely to remain active even after exposure to USP lasers. The viruses that were tested for inactivation by USP laser in human plasma were an enveloped RNA virus human immunodeficiency virus (HIV), nonenveloped RNA virus hepatitis A virus (HAV) and enveloped DNA virus murine cytomegalovirus (MCMV). Continue reading “Using Laser Treatment to Eliminate Blood-Borne Pathogens”
When DNA Is Not Enough: New Research Suggests Epigenetic Factors Play an Important Role in the High Mortality Rate of the Devi Facial Tumor Disease
 The Devil Facial Tumor Disease (DFTD) is a contagious cancer in Tasmanian Devils that is threatening the species with extinction. This disease is spread from individual to individual and has a 100% mortality rate. It is so deadly because, although the DFTF cells should be attached and killed by the host devil’s immune system, for some reason they are not—and no one is sure why. A study published in PNAS in March of last year (1) showed that DFTD cells don’t express surface MHC molecules. MHC class I and class II molecules are crucial for proper immune response, and their absence on the cell surface could explain why the DFTD cells do not stimulate an immune response.
The Devil Facial Tumor Disease (DFTD) is a contagious cancer in Tasmanian Devils that is threatening the species with extinction. This disease is spread from individual to individual and has a 100% mortality rate. It is so deadly because, although the DFTF cells should be attached and killed by the host devil’s immune system, for some reason they are not—and no one is sure why. A study published in PNAS in March of last year (1) showed that DFTD cells don’t express surface MHC molecules. MHC class I and class II molecules are crucial for proper immune response, and their absence on the cell surface could explain why the DFTD cells do not stimulate an immune response.
The authors found that the loss of MHC expression is maintained as the cells divide, and is not a result of structural mutations in the genes responsible for MHC expression. Instead the authors found that this down regulation was the result of regulatory changes including epigenetic modifications to histones. Continue reading “When DNA Is Not Enough: New Research Suggests Epigenetic Factors Play an Important Role in the High Mortality Rate of the Devi Facial Tumor Disease”
Mass Spectrometry Application: Antibody Quantitation for Preclinical PK studies
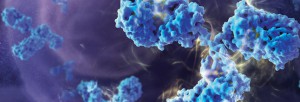 Therapeutic monoclonal antibodies (mAbs) represent the majority of therapeutics biologics now on the market, with more than 20 mAbs approved as drugs (1–3). During preclinical development of therapeutic antibodies, multiple variants of each antibody are assessed for pharmacokinetic (PK) characteristics across model systems such as rodents, beagles and primates. Ligand-binding assays (LBA) are the standard technology used to perform the PK studies for mAb candidates (4). Ligand-binding assays (LBAs) are methods used to detect and measure a macromolecular interaction between a ligand and a binding molecule. In LBAs, a therapeutic monoclonal antibody is considered to be the ligand, or analyte of interest, while the binding molecule is usually a target protein.
Therapeutic monoclonal antibodies (mAbs) represent the majority of therapeutics biologics now on the market, with more than 20 mAbs approved as drugs (1–3). During preclinical development of therapeutic antibodies, multiple variants of each antibody are assessed for pharmacokinetic (PK) characteristics across model systems such as rodents, beagles and primates. Ligand-binding assays (LBA) are the standard technology used to perform the PK studies for mAb candidates (4). Ligand-binding assays (LBAs) are methods used to detect and measure a macromolecular interaction between a ligand and a binding molecule. In LBAs, a therapeutic monoclonal antibody is considered to be the ligand, or analyte of interest, while the binding molecule is usually a target protein.
LBAs have certain well-documented limitations (5). Specific assay reagents are often not available early in a program. Interferences from endogenous proteins, antidrug antibodies, and soluble target ligands are potential complicating factors.
Liquid chromatography coupled to tandem mass spectrometry (LC–MS/MS)-based methods represent a viable and complementary addition to LBA for mAb quantification in biological matrixes. LC–MS/MS provides specificity, sensitivity, and multiplexing capability.
A recent reference (6) illustrates an automated method to perform LC–MS/MS-based quantitation, with IgG1 conserved peptides, a heavy isotope labeled mAb internal standard,and anti-human Fc enrichment. The method was applied to the pharmacokinetic study of a mAb dosed in cynomolgus monkey, and the results were compared with the immunoassay data. The interesting finding of the difference between ELISA and LC–MRM-MS data indicated that those two methods can provide complementary information regarding the drug’s PK profile.
Literature Cited
- Mao, T. et al. (2013) Top-Down Structural Analysis of an Intact Monoclonal Antibody by Electron Capture Dissociation-Fourier Transform Ion Cyclotron Resonance-Mass Spectrometry. Anal.Chem. 85, 4239–46.
- Weiner, L. M. et al. (2010) Monoclonal antibodies: versatile platforms for cancer immunotherapy. Nat. Rev. Immunol. 10, 317–27.
- Nelson, A. et al. (2010) Development trends for human monoclonal antibody therapeutics. Nat. Rev. Drug Discovery. 9, 767–74.
- DeSilva, B. et al. (2003) Recommendations for the Bioanalytical Method Validation of Ligand-Binding Assays to Support Pharmacokinetic Assessments of Macromolecules. Pharm. Res. 20, 1885–00.
- Ezan, E.et al. (2009) Critical comparison of MS and immunoassays for the bioanalysis of therapeutic antibodies. Bioanalysis 1, 1375–88.
- Zhang, Q. et al. (2014) Generic Automated Method for Liquid Chromatography–Multiple Reaction Monitoring Mass Spectrometry Based Monoclonal Antibody Quantitation for Preclinical Pharmacokinetic Studies. Anal.Chem. 86, 8776–84.
Is Artificial Intelligence a Threat to Mankind?
 Technology: We all use it, and some of us couldn’t go an entire day without it. In many ways, digital technology has improved our lives by increasing productivity and communication. Computer technology is everywhere: our homes, offices, phones and even cars. Technology has integrating into our lives so completely that most of us no longer stop to marvel at even the [seemingly] simplest capabilities such as the predictive software that our smart phones use to predict which word we are typing after we type in only the first few letters, especially if the software gets it wrong much of the time. However, digital technology has its dangers and inconveniences: cybercrime, hackers, stolen data, and computer crashes and failed Wi-Fi connections at the most inopportune times. In a recent BBC interview, one of modern science’s most brilliant minds highlighted another potential danger: artificial intelligence. Does artificial intelligence pose a threat to mankind?
Technology: We all use it, and some of us couldn’t go an entire day without it. In many ways, digital technology has improved our lives by increasing productivity and communication. Computer technology is everywhere: our homes, offices, phones and even cars. Technology has integrating into our lives so completely that most of us no longer stop to marvel at even the [seemingly] simplest capabilities such as the predictive software that our smart phones use to predict which word we are typing after we type in only the first few letters, especially if the software gets it wrong much of the time. However, digital technology has its dangers and inconveniences: cybercrime, hackers, stolen data, and computer crashes and failed Wi-Fi connections at the most inopportune times. In a recent BBC interview, one of modern science’s most brilliant minds highlighted another potential danger: artificial intelligence. Does artificial intelligence pose a threat to mankind?
Continue reading “Is Artificial Intelligence a Threat to Mankind?”
Christensenellaceae—A Natural Way to Stay Thin?
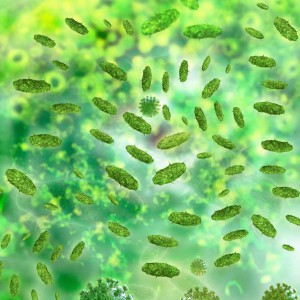 A study published in the Nov 6 issue of Cell outlined results suggesting that an obscure family of bacteria colonizing the human gut may be inherited and may also have a direct influence on body weight. The paper is the first to identify such an association and to link a particular microbial colonist with lower BMI. Continue reading “Christensenellaceae—A Natural Way to Stay Thin?”
A study published in the Nov 6 issue of Cell outlined results suggesting that an obscure family of bacteria colonizing the human gut may be inherited and may also have a direct influence on body weight. The paper is the first to identify such an association and to link a particular microbial colonist with lower BMI. Continue reading “Christensenellaceae—A Natural Way to Stay Thin?”
Career Development for Working Professionals: UW Master of Science in Biotechnology Program
Prior to the Masters in Biotechnology program, I had no working knowledge of Intellectual Property (IP), e.g., patents, trademarks, etc. The M.S. in Biotechnology program not only opened my eyes to Intellectual Property and its importance in biotechnology companies, but it sparked my interested in a career in an IP field. From the knowledge I gained and connections made in the program, I have been able to achieve a career in IP. I am now happy to be able to share my experience and knowledge with current and future students in the program.
—Heather Gerard, M.S. (2006) Intellectual Property Manager, Promega Corporation
Since 2002, the BioPharmaceutical Technology Center Institute (BTC Institute) has been effectively collaborating with the UW Master of Science in Biotechnology Program (MS-Biotechnology) to provide the three lab-based Molecular Technologies courses for this unique degree designed for working professionals.
As noted on the program’s web site, it offers:
- A curriculum like no other that integrates topics in science, business and law
- Powerful skills that bring the “big picture” of life science product development into clear focus
- Exclusive evening/weekend courses allowing you to work full-time while enrolled, and
- A completed degree in less than two years
What Things Are You Thankful for in Science?

I wondered what “thanksgiving” looks like to the research scientist. So I asked:
What are the things you are thankful for in science?
The answers have been as varied as the people I talked to ranging from little things like water bath floats to really big things, like the renewal of your research funding or achieving tenure.
Here are some of the answers from my informal inquiries:
“Tube floaties for water baths.”
—E.V., genomics product manager
“I was always thankful for Geiger counters.”
—K. G., science writer
“Thermal cyclers and Taq Polymerase. As an undergrad I watched someone sit with a timer and move their tubes between water baths at 3 different temperatures, opening tubes and adding polymerase at the end of each cycle. Modern PCR is SOOO much easier.”
—M.M., research scientist
“I am thankful for competent cells. I remember preparing the CaCl2 and doing slow centrifugation. Also thankful for serum-compatible transfection, rapid ligations and online journal access (no longer have to traipse over to the university library to get papers photocopied- uuurrrgggghhh).”
—R.D., technical services scientist
“How about T-vectors for cloning? I was no molecular biologist, but could make a T-vector work.”
—K.K., science writer
“I am thankful for open-access journals and the ability to read the full article without an institutional subscription.”
—S.K., science writer
“I am ever so thankful for ONLINE ORDERING! So awesome. Throw in online technical manuals, on-line support tools, on-line calculators – all are awesome!!”
—A.P., director, scientific courses
“I am thankful for automated sequencing- manual sequencing was laborious and hazardous!!!”
—R.G., technical services scientist
Do any of these resonate with you? What are you thankful for as a scientist? Let us know in the comments.
DNA Amplification Using Body Heat, No Instrument Required

Insights into the Function of P7C3 Compounds in Neuroprotection

It is fall and the season for American football. For this football fan, watching the game is a bit less enjoyable than it used to be, as more and more information is available about the serious and permanent brain injuries suffered by football players.
In the introduction to a recent paper in the journal Cell, “P7C3 Neuroprotective Chemicals Function by Activating the Rate-Limiting Enzyme in NAD Salvage”, not a word about American football is mentioned.
However, the paper begins, “No substantive therapeutics are available for the treatment of almost any form of disease entailing nerve death” (1). The authors list a range of neurodegenerative disorders such as Huntington’s, Alzheimers and Parkinson’s diseases, as well as ALS or Lou Gherig’s disease. They also note that there are currently no effective treatments for trauma to the brain or peripheral nervous system.
The authors note that a chemical treatment that could interfere with nerve cell death would have a “transformative impact in modern medicine”. Continue reading “Insights into the Function of P7C3 Compounds in Neuroprotection”
Optimizing Tryptic Digestions for Phosphoproteomics Analysis
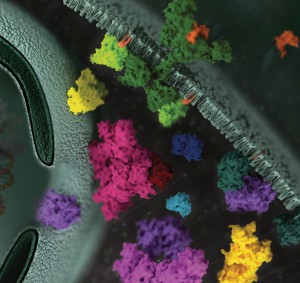 Protein phosphorylation is the most widespread type of post-translational modification. It affects every basic cellular process, including metabolism, growth, division, differentiation, motility, organelle trafficking, membrane transport, muscle contraction, immunity, learning and memory (1,2). Protein kinases catalyse the transfer of the phosphate from ATP to specific amino acids in proteins. In eukaryotes, these are usually Ser, Thr and Tyr residues. Due to the development of specific phosphopeptide enrichment techniques and highly sensitive MS instruments, phosphoproteomics has enabled researchers to gain a comprehensive view on the dynamics of protein phosphorylation and phosphorylation based signaling networks.
Protein phosphorylation is the most widespread type of post-translational modification. It affects every basic cellular process, including metabolism, growth, division, differentiation, motility, organelle trafficking, membrane transport, muscle contraction, immunity, learning and memory (1,2). Protein kinases catalyse the transfer of the phosphate from ATP to specific amino acids in proteins. In eukaryotes, these are usually Ser, Thr and Tyr residues. Due to the development of specific phosphopeptide enrichment techniques and highly sensitive MS instruments, phosphoproteomics has enabled researchers to gain a comprehensive view on the dynamics of protein phosphorylation and phosphorylation based signaling networks.
Due to its high cleavage specificity, trypsin is the commonly used proteolytic enzyme in MS-based proteomics, cleaving peptides carboxyterminal of the amino acids lysine and arginine. However, various factors such as the tertiary structure of a protein, adjacent basic amino acids or negatively charged residues close to cleavage sites as well as PTMs are known to impair proteolysis.
To gain closer insights into the impact of phosphorylation on tryptic digestion, a recent publication(3) systematically characterized the digestion efficiency of model peptide sequences that are known to be prone to incomplete digestion.
The results indicated that increasing trypsin concentrations up to a trypsin to peptide ratio of 1:10 led to a significant gain (1) in the overall number of phosphorylation sites (up to 9%) and in the intensities of individual phosphopeptides, thereby improving the sensitivity of phosphopeptide quantification.
The effect of organic solvents (ACN, acetonitrile and TFE trifuorethanol was also evaluated). Positive results were noted with TFE when determining the digestion of individual peptides. However TFE interfered with TiO2 phosphopeptide enrichment and therefore was not recommended for use with complex samples.
- Engholm-Keller, K and Larsen, M.R. (2013) Technologies and challenges in large scale phosphoroproteomics. Proteomics 13, 910–31.
- Beausoleil, S. A. et al. (2010) Tissue-specific atlas of mouse protein phosphorylation and expression. Cell 143, 1174–89.
- Dickhut, C. et al. (2014) Impact of Digestion Conditions on phosphoproteomics. J. Proteome Res. 13, 2761–70.
