 Filter-aided sample preparation (FASP) method is used for the on-filter digestion of proteins prior to mass-spectrometry-based analyses (1,2). FASP was designed for the removal of detergents, and chaotropes that were used for sample preparation. In addition, FASP removes components such as salts, nucleic acids and lipids. Akylation of reduced cysteine residues is also carried out on filter, after which protein is proteolyzed by use of trypsin on filter in the optimal buffer of the enzyme. Subsequent elution and desalting of the peptide-rich solution then provides a sample ready for LC–MS/MS analysis.
Filter-aided sample preparation (FASP) method is used for the on-filter digestion of proteins prior to mass-spectrometry-based analyses (1,2). FASP was designed for the removal of detergents, and chaotropes that were used for sample preparation. In addition, FASP removes components such as salts, nucleic acids and lipids. Akylation of reduced cysteine residues is also carried out on filter, after which protein is proteolyzed by use of trypsin on filter in the optimal buffer of the enzyme. Subsequent elution and desalting of the peptide-rich solution then provides a sample ready for LC–MS/MS analysis.
Erde et al. (3) described an enhanced FASP (eFASP) workflow that included 0.2% DCA in the exchange, alkylation, and digestion buffers,thus enhancing trypsin proteolysis, resulting in increases cytosolic and membrane protein representation. DCA has been reported (4) to improve the efficiency of the denaturation, solubilization, and tryptic digestion of proteins, particularly proteolytically resistant myoglobin and integral membrane proteins, thereby enhancing the efficiency of their identification with regard to the number of identified proteins and unique peptides.
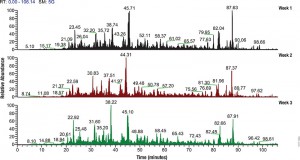
In a recent publication (5) traditional FASP and eFASP were re-evaluated by ultra-high-performance liquid chromatography coupled to a quadrupole mass filter Orbitrap analyzer (Q Exactive). The results indicate that at the protein level, both methods extracted essentially the same number of hydrophobic transmembrane containing proteins as well as proteins associated with the cytoplasm or the cytoplasmic and outer membranes.
The LC–MS/MS results indicate that FASP and eFASP showed no significant differences at the protein level. However, because of the slight differences in selectivity at the physicochemical level of peptides, these methods can be seen to be somewhat complementary for analyses of complex peptide mixtures.
- Manza, L. L. et al. (2005) Sample preparation and digestion for proteomic analyses using spin filters Proteomics 5, 1742–74.
- Wiśniewski, J. R. et al. (2009) Universal sample preparation method for proteome analysis Nat. Methods 6, 359–62.
- Erde, J. et al. (2014) Enhanced FASP (eFASP) to increase proteomic coverage and sample recovery for quantitative proteome experiments. J. Proteome Res. 13, 1885–95.
- Lin, Y. et al. (2008) Sodium-deoxycholate-assisted tryptic digestion and identification of proteolytically resistant proteins Anal. Biochem. 377, 259–66.
- Nel. A. et al. (2015) Comparative Reevaluation of FASP and Enhanced FASP methods by LC-MS/MS/ J Proteome Res. 14, 1637–42.
