Huang et al. were interested in assessing both mutations and gene expression levels in colorectal cancer (CRC) using a single detection method. They developed an analysis they called multiplex ligation-dependent probe amplification–digital amplification coupled with hydrogel bead-array (MLPA–DABA). This method involves several probes that hybridize to the target sequence and form a template for PCR only if the left probe and the right probe were both completely complementary with the target. If there was a single base pair on the 5′ end of the right probe that does not hybridize, ligation of the left and right probes into a single template used in the later PCR analysis did not occur. To detect mutations, the left probe consisted of a universal primer sequence and a gene-specific sequence. The two right probes had a gene-specific sequence that reflected either wildtype or mutant sequence with the base difference at the 5′ end, a sequence for Cy®3 or Cy®5 probe binding and a universal primer sequence. For gene expression analysis, the left and right probes were similar to the mutation detection left and right probes with the exception that there was only one gene-specific sequence for the right probe. The Cy®5 sequence was attached to wildtype and housekeeping genes; Cy®3 sequence attached to mutant and colorectal cancer-related genes.
Once the DNA or RNA was mixed with the gene expression or mutation probes and ligation occurred, the next step was emulsion PCR. This involved an oil and aqueous mixture with beads that had a PCR primer immobilized on it. The oil with an amplification mix (the aqueous phase) was mixed to get the smallest droplets for single-molecule PCR (digital PCR). By adding the ligated template and primers with additional amplification mix, the template and beads are diluted to the point that a single template and a single bead are together in one aqueous droplet and PCR proceeds in the droplet, amplifying only the one template. After amplification, the beds are washed in isopropanol to remove the emulsification oil, the beads filtered and washed, the DNA treated with NaOH to created ssDNA immobilized on beads before creating a single bead array on a slide. Free and mismatched probes are removed from the array using electrophoresis before Cy®3 and Cy®5 probes are added for detection.
To test this method, Huang et al. used β-actin and tagged it in the MLPA portion with the Cy®5 tag sequence. When β-actin was amplified, one of the primers was labeled with biotin and immobilized on streptavidin. The subsequent array was incubated with both Cy®3 and Cy®5 but only the red Cy®5 fluoresced on the bead array. Conversely, the cDNA from CRC genes was only labeled with Cy®3 tag sequence for gene expression analysis, and after incubation with both Cy®3 and Cy®5, only fluoresced green. Researchers were confident that the electrophoresis step removed mismatched or residual probes from the hydrogel bead array. In addition, the number of falsely scored beads for both experiments were nearly zero, indicating the assay is specific for the genes being analyzed. When performing mutational analysis of SW480 cells with a known homozygous mutation at codon 1338, scientists found the same nearly zero falsely scored beads.
Quantitation of two genes to test the MLPA–DABA method was done comparing two housekeeping genes: β-actin and GAPDH. β-actin was labeled with Cy®5 sequence and GAPDH labeled with Cy®3 sequence tag. The number of red and green beads were basically 1:1, a successful detection of these two genes. Furthermore, none of the beads fluoresced yellow, which would suggest both genes were amplified in a single droplet prior to detection.
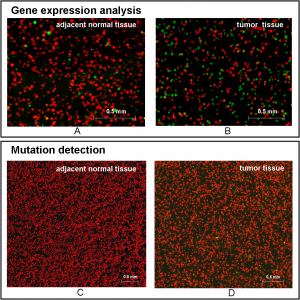
So how does these technique work for detecting CRC gene expression and mutations? Using both tumor and normal tissue from individuals, β-actin plus five CRC-related genes (c-Myc, h-Ras, CD44v6, Cox-2 and n-Ras) were assessed for expression levels. For tumor tissue, the expression levels of the five genes were elevated versus normal tissue when compared the level of β-actin. The mutational analysis demonstrated no mutations in the normal tissue, but there were mutations in the tumor tissue. Based on the ratio of red (wildtype) to green (mutation), the mutation rate was calculated at 3.1%.
To extend the work into a noninvasive testing method, DNA was isolated from stool samples of individuals with CRC. For the six samples tested, 3 of the 6 individuals (50%) had mutations while five of the six had increased gene expressions levels of the five CRC-related genes. These results suggest that the MLPA–DABA is a promising assay, especially for early stage detection. Four of the samples came from individuals with Dukes stage A and B (the earlier two of a four-stage classification) and MLPA-DABA method successfully detected changes in three of the samples.
The method developed by Huang et al. is a promising assay for detecting colorectal cancer and may even prove to be useful for noninvasive detection of early stages of CRC. The panel of genes used in the PLOS ONE article is limited so increasing utility and discrimination of the assay would mean analyzing more CRC-specific genes.
Reference
Huang, H., Li, S., Sun, L. and Zhou, G. (2015) Digital detection of multiple minority mutants and expression levels of multiple colorectal cancer-related genes using digital-PCR coupled with bead-array. PLOS ONE 10, e0123420.
Sara Klink
Latest posts by Sara Klink (see all)
- A One-Two Punch to Knock Out HIV - September 28, 2021
- Toxicity Studies in Organoid Models: Developing an Alternative to Animal Testing - June 10, 2021
- Herd Immunity: What the Flock Are You Talking About? - May 10, 2021
