Bottom-up proteomics focuses on the analysis of protein mixtures after enzymatic digestion of the proteins into peptides. The resulting complex mixture of peptides is analyzed by reverse-phase liquid chromatography (RP-LC) coupled to tandem mass spectrometry (MS/MS). Identification of peptides and subsequently proteins is completed by matching peptide fragment ion spectra to theoretical spectra generated from protein databases.
Trypsin has become the gold standard for protein digestion to peptides for shotgun proteomics. Trypsin is a serine protease. It cleaves proteins into peptides with an average size of 700-1500 daltons, which is in the ideal range for MS (1). It is highly specific, cutting at the carboxyl side of arginine and lysine residues. The C-terminal arginine and lysine peptides are charged, making them detectable by MS. Trypsin is highly active and tolerant of many additives.
Even with these technical features, the use of trypsin in bottom-up proteomics may impose certain limits in the ability to grasp the full proteome, Tightly-folded proteins can resist trypsin digestion. Post-translational modifications (PTMs) present a different challenge for trypsin because glycans often limit trypsin access to cleavage sites, and acetylation makes lysine and arginine residues resistant to trypsin digestion.
Are you looking for proteases to use in your research?
Explore our portfolio of proteases today.
To overcome these problems, the proteomics community has begun to explore alternative proteases to complement trypsin. However, protocols, as well as expected results generated when using these alternative proteases have not been systematically documented.
In a recent reference (2), optimized protocols for six alternative proteases that have already shown promise in their applicability in proteomics, namely chymotrypsin, Lys-C, Lys-N, Asp-N, Glu-C and Arg-C have been created.
Data describe the appropriate MS data analysis methods and the anticipated results in the case of the analysis of a single protein (BSA) and a more complex cellular lysate (Escherichia coli). The digestion protocol presented here is convenient and robust and can be completed in approximately in 2 days.
Try a sample of high-efficiency Trypsin Platinum today!
Visit our website for more on Trypsin Platinum, Mass Spectrometry Grade, with enhanced proteolytic efficiency and superior autoproteolytic resistance.
Reference
- Laskay, U. et al. (2013) Proteome Digestion Specificity Analysis for the Rational Design of Extended Bottom-up and middle-down proteomics experiments. J of Proteome Res. 12, 5558–69.
- Giansanti, P. et. al. (2016) Six alternative protease for mass spectrometry based proteomics beyond trypsin. Nat. Protocols 11, 993–6
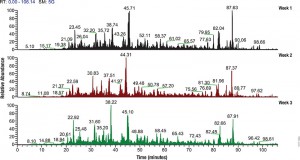 Filter-aided sample preparation (FASP) method is used for the on-filter digestion of proteins prior to mass-spectrometry-based analyses (1,2). FASP was designed for the removal of detergents, and chaotropes that were used for sample preparation. In addition, FASP removes components such as salts, nucleic acids and lipids. Akylation of reduced cysteine residues is also carried out on filter, after which protein is proteolyzed by use of trypsin on filter in the optimal buffer of the enzyme. Subsequent elution and desalting of the peptide-rich solution then provides a sample ready for LC–MS/MS analysis.
Filter-aided sample preparation (FASP) method is used for the on-filter digestion of proteins prior to mass-spectrometry-based analyses (1,2). FASP was designed for the removal of detergents, and chaotropes that were used for sample preparation. In addition, FASP removes components such as salts, nucleic acids and lipids. Akylation of reduced cysteine residues is also carried out on filter, after which protein is proteolyzed by use of trypsin on filter in the optimal buffer of the enzyme. Subsequent elution and desalting of the peptide-rich solution then provides a sample ready for LC–MS/MS analysis.