The Medicinal Chemistry Center (CQMED), headquartered at Campinas State University in Brazil, recently started a project in partnership with Promega to develop drugs that can be used against Leishmania. This genus of protozoans is the etiological agent of leishmaniasis, transmitted to humans by sandflies.
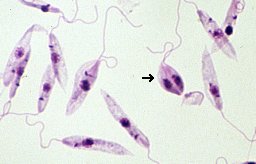
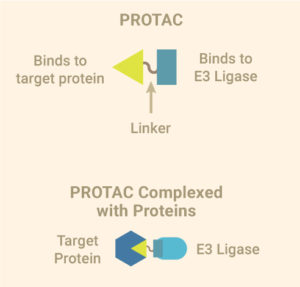
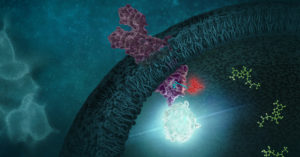
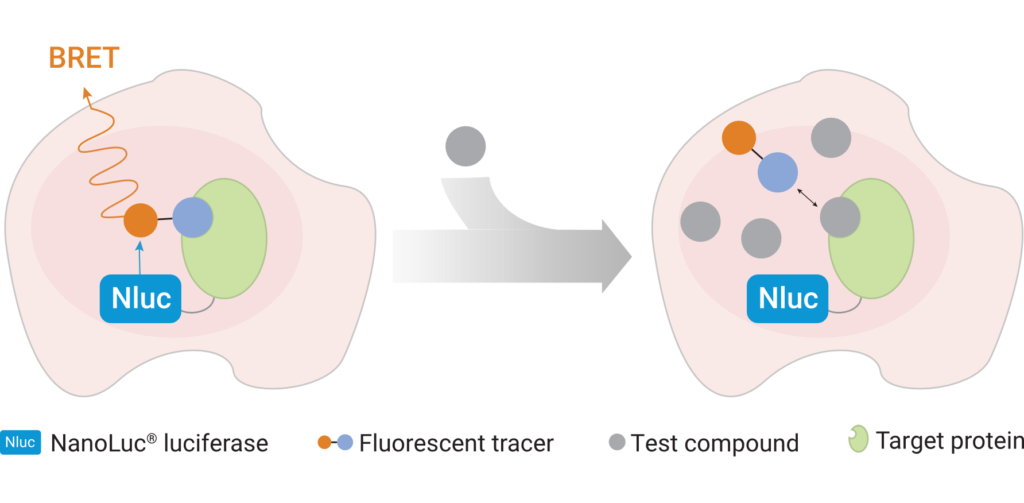
Leishmaniasis is classified as a neglected tropical disease that mainly affects poor communities. Symptoms include large skin sores and an enlarged spleen. The challenge in developing drugs to treat Leishmania is finding appropriate therapeutic targets. These targets are normally proteins whose inhibition leads to death of the parasite. In addition to pharmaceutical company Eurofarma, whose goal is to develop drugs for Leishmania, Promega was chosen to help solve this problem because of our NanoBRET™ Target Engagement (TE) assay*, a well-established technique for measuring protein interactions. In this assay, NanoLuc® luciferase is attached to the protein of interest, and a fluorescent NanoBRET™ tracer molecule is added to the cells. This produces a BRET signal. When a competing ligand is added, it will displace the tracer molecule, enabling quantification of the strength of the interaction compared to the tracer molecule..
A challenge that researchers will face will be ensuring that the NanoBRET™ tracer reaches the inside of the parasite cells; because Leishmania is an intracellular parasite, molecules need to cross the host cell membrane, the membrane of the vacuole containing the parasites, and the membrane of the parasite itself. Another challenge the slow reproduction of Leishmania within macrophages. On top of that is the fact that the parasite’s metabolism varies depending on its biological cycle, meaning that there could be long periods of time during which a drug’s therapeutic target is not expressed in the cell, during which time the drug would have no effect. The ideal target would be expressed at high levels throughout the cell cycle.
The project is being led by Rafael Couñago, a researcher at CQMED, and Promega scientists Matt Robers and Jean-Luc Vaillaud.
*An earlier version of this blog incorrectly said that these experiments are based on the NanoBRET™ assay using HaloTag® protein.