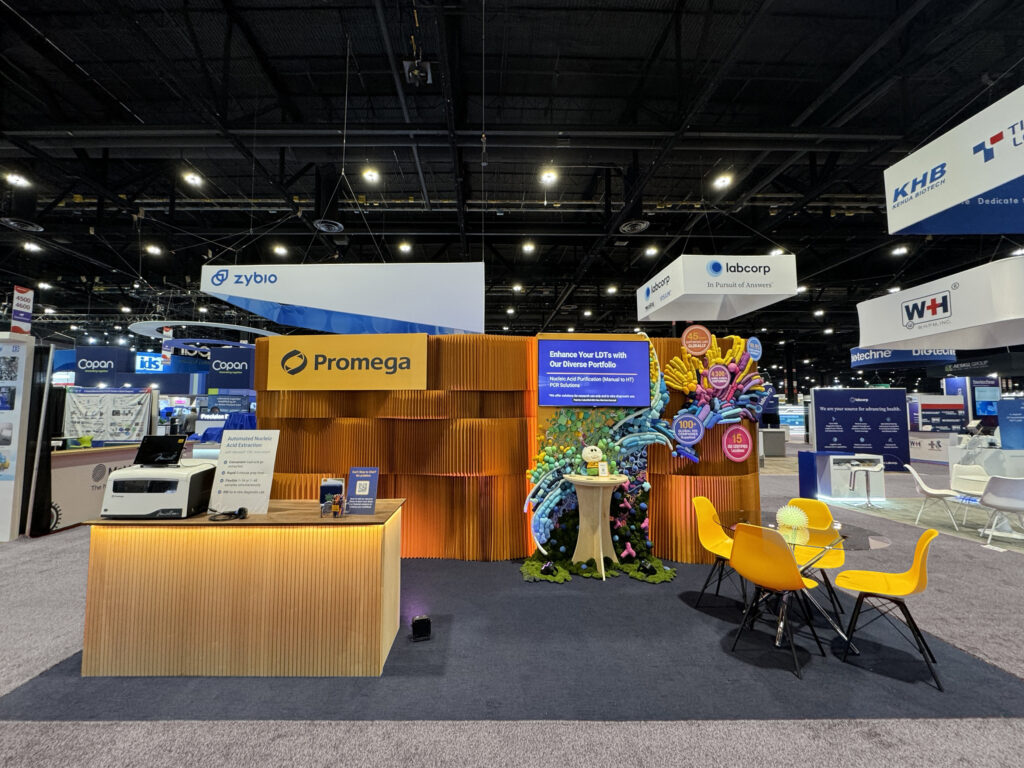
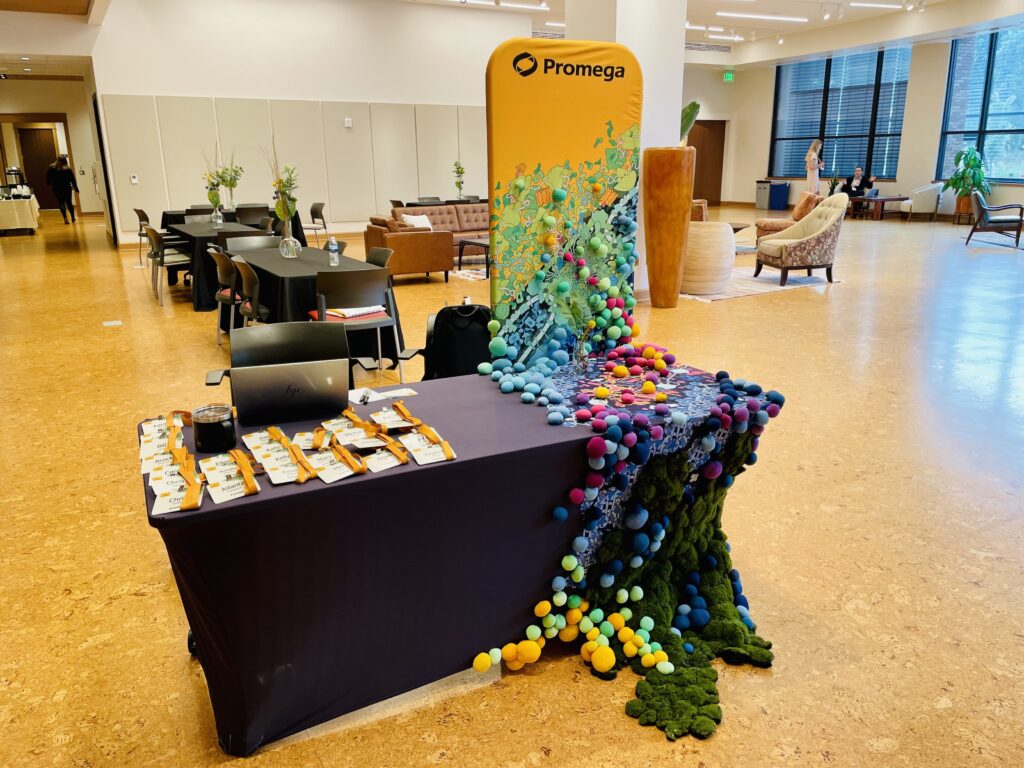
Step inside a Promega booth and leave the ordinary behind. Here, science sparks creativity, sustainability is woven into every detail, and discovery isn’t just something you see—it’s something you feel.
Continue reading “Bringing Science to Life: How Art and Sustainability Shape Our New Trade Show Booth Design”#PromegaWorks
Descriptions of Promega products in action from examples of Promega products in the peer-reviewed literature to technical tips on how to address common laboratory and experimental questions
Decades of Discovery: How the NCI-60 Revolutionized Cancer Drug Screening

The National Cancer Institute’s NCI-60 drug screening panel, comprised of 60 diverse human cancer cell lines, has been a cornerstone in advancing cancer research and drug discovery since its inception in the late 1980s. Developed in response to the need for more predictive and comprehensive preclinical models, the NCI-60 facilitates the screening of thousands of compounds annually, aiming to identify potential anti-cancer drugs across a broad spectrum of human cancers. This article traces the origins, development, and evolution of the NCI-60 panel, highlighting its significant role in advancing our understanding of cancer and therapeutic agents.
Continue reading “Decades of Discovery: How the NCI-60 Revolutionized Cancer Drug Screening”Raising Frogs Takes a Village: Accelerating Amphibian Research at the Marine Biological Laboratory
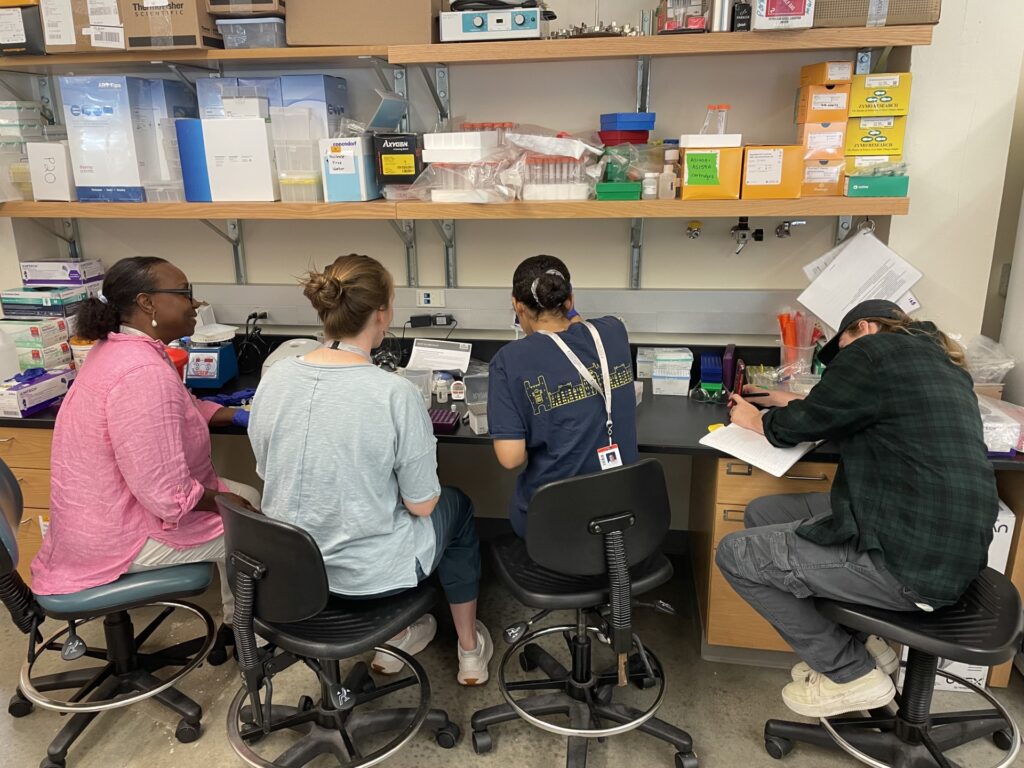

Sally Seraphin’s life in the research lab started with rats and roseate terns. Chimpanzees and rhesus macaques came next, then humans (and a brief foray into voles). When she pivoted to red-eyed tree frogs, Sally once again had to learn all kinds of new techniques. Suddenly, in addition to new sample prep and analysis techniques, she needed to get up to speed on amphibian care and husbandry. That led her to the Marine Biological Laboratory (MBL) in Woods Hole, MA.
“It’s a seaside resort atmosphere with experts in every technology you can imagine,” Sally says. “It’s a place to incubate and birth new approaches to answering questions.”
Sally spent the past two summers at MBL learning everything she needed to know about breeding and caring for amphibians. During that time, she also worked closely with Applications Scientists from Promega who helped her start extracting RNA from frog samples.
“The hands-on support from industry scientists is definitely unique to Promega and MBL,” she says. “It’s rare to have a specialist on hand who can help you learn, troubleshoot and optimize in such a finite amount of time.”
Adopting a New Model Organism
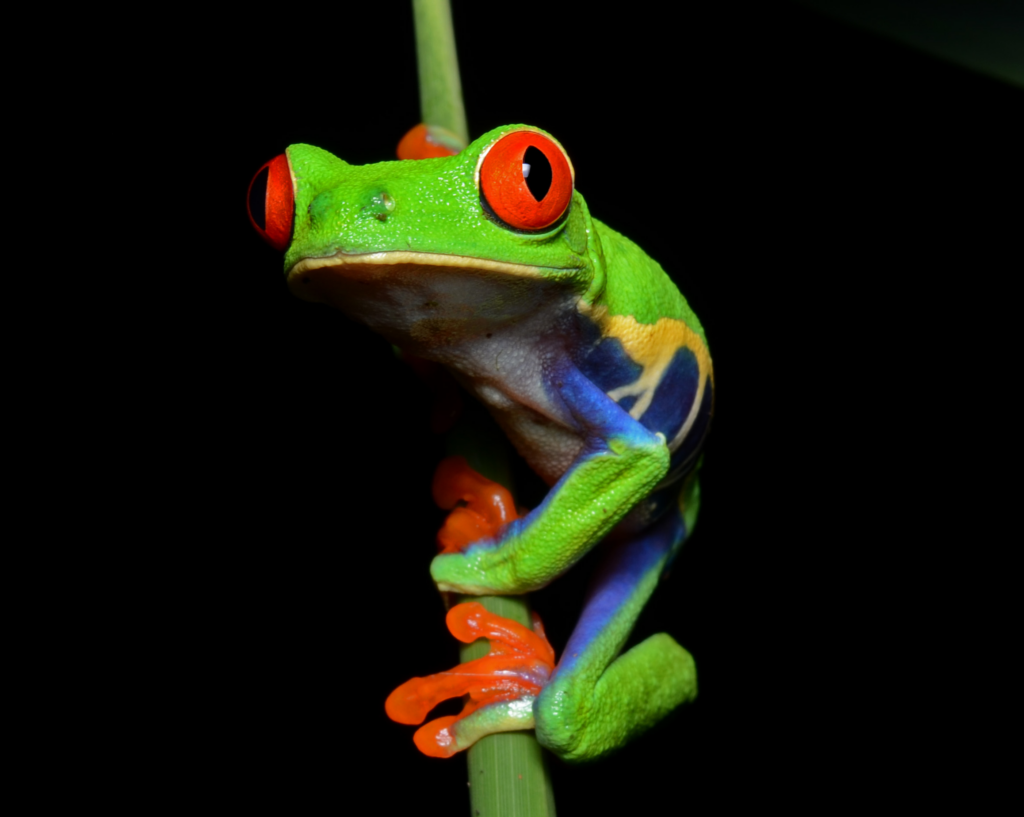

Sally studies how early stress impacts brain and behavior development. She hopes to deepen our understanding of how adverse childhood experiences connect to mental illness and bodily disease later in life. In the past, she studied how factors such as parental absence affected the neurotransmission of dopamine in primates. Recently, she changed her focus to developmental timing.
“Girls who are exposed to early trauma like sexual or physical abuse will sometimes reach puberty earlier than girls who aren’t,” Sally explains. “And I noticed that there are many species that will alter their developmental timing in response to predators or social and ecological threats.”
Continue reading “Raising Frogs Takes a Village: Accelerating Amphibian Research at the Marine Biological Laboratory”Uniting Diverse Minds, Vibrant Ideas, and Collaborative Spirit at the 2023 Biologics Symposium
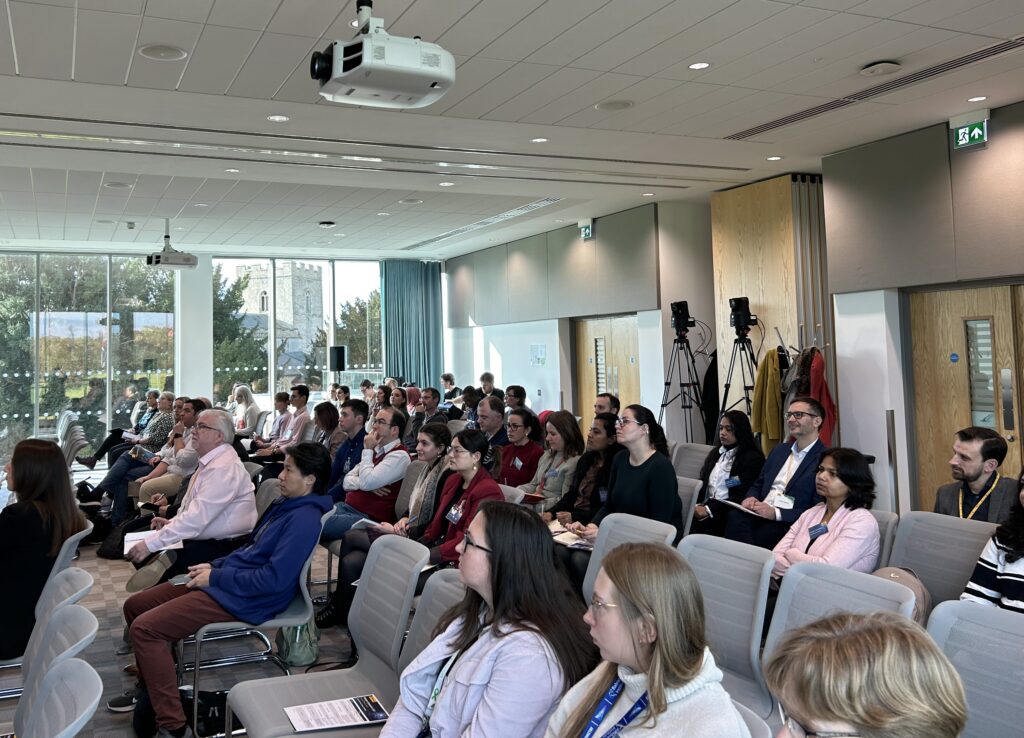

On Thursday November 9th, 2023, Promega held its 7th Biologics Symposium at the Babraham Research Campus in Cambridge. For the first time, participants had the option to attend the event either in person or experience it via live stream, creating an inclusive and dynamic environment where the latest breakthroughs and ideas could be showcased. Moreover, the event was organized into a morning and afternoon session, enabling ample time for networking and the exchange of ideas beyond formal presentations.
Continue reading “Uniting Diverse Minds, Vibrant Ideas, and Collaborative Spirit at the 2023 Biologics Symposium”Custom in vitro Transcription Reagents for Manufacturing RNA Therapeutics
Research into vaccines based on RNA began decades ago when scientists theorized that they could harness RNA to produce viral proteins within a cell, prompting a protective immune response. RNA vaccine research drew scientists’ attention during the development of SARS-CoV-2 vaccines during the COVID-19 pandemic, which opened the door for research targeting other diseases with RNA-based therapeutics.
Continue reading “Custom in vitro Transcription Reagents for Manufacturing RNA Therapeutics”We’re Committing to 100% Renewable Electricity by 2025
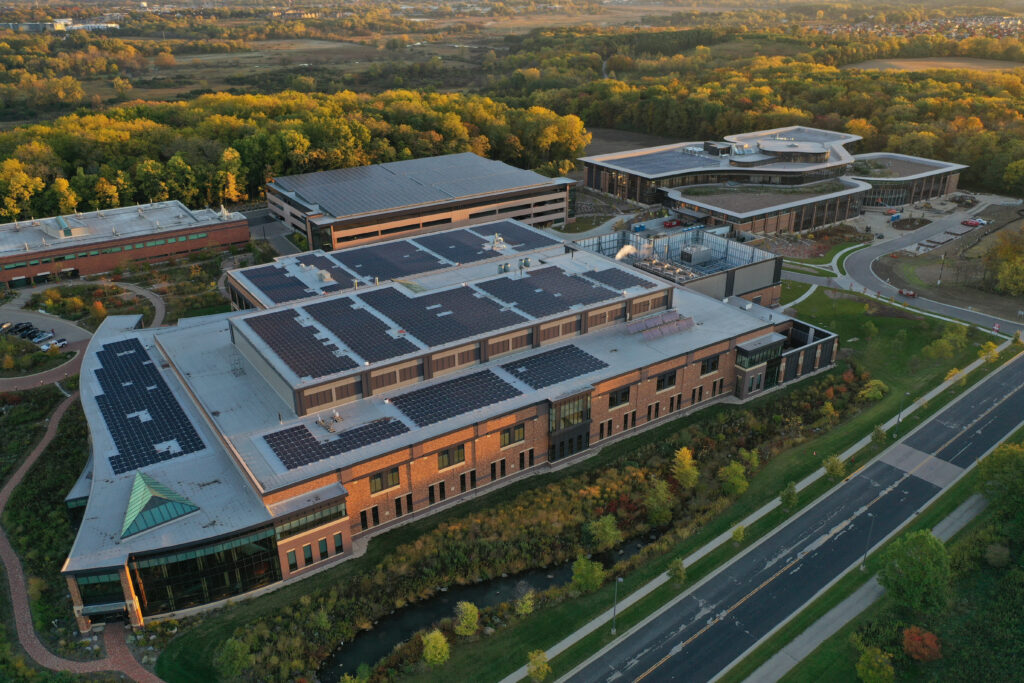

In 2021, we unveiled our most ambitious sustainability goals ever. These goals include a 50% reduction in carbon emissions by 2030, as indexed to revenue over a 2019 baseline.
In 2022, we announced that renewable sources provided over 20% of our global electricity usage.
This year, Promega is excited to announce that we’re committing to 100% renewable electricity by 2025.
Continue reading “We’re Committing to 100% Renewable Electricity by 2025”Custom Manufacturing: Translating Research into Product
Scientists around the world are focusing their energy and resources on translating advances made in clinical research into relevant biotechnology, clinical, and applied products that improve our health and well-being. Once research looks promising, there is substantial pressure to expedite the release of that product or assay in the market.
For many organizations focused on developing these advanced products, their expertise and core competencies are in developing the assay. Often, they do not have the experience, infrastructure, or quality systems in place to support large-scale production, packaging, or distribution of their newly developed assay in a way that is also in compliance with relevant regulatory requirements. These next steps become a barrier to realizing the value of the research. Working with a custom or contract manufacturing partner can lower this barrier and expedite the time to market.
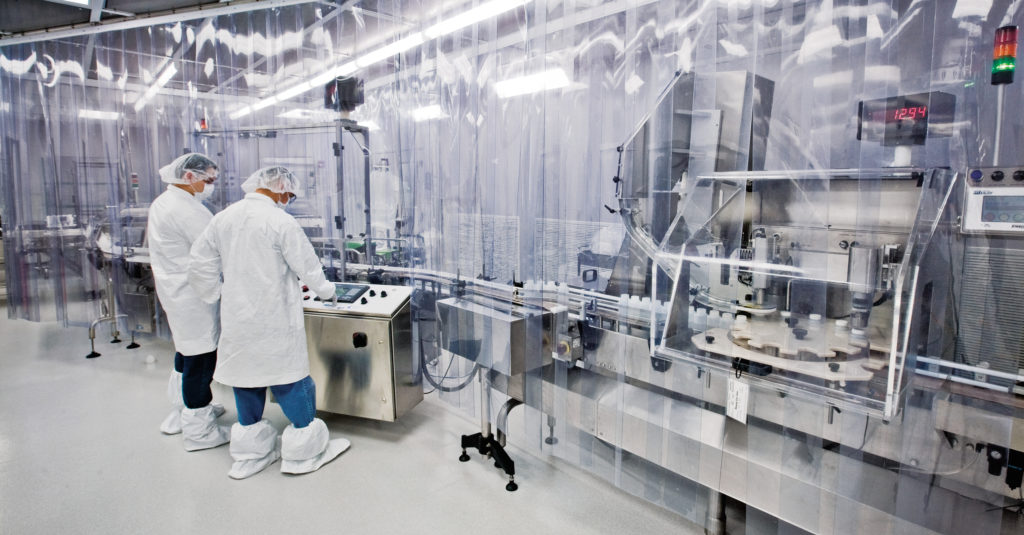

Be careful not to confuse custom manufacturing with original equipment manufacturer (OEM) products. OEM products are existing products from one company that another company rebrands and sells. Custom manufacturers typically focus on providing more comprehensive services that can be adapted to produce a new product. Custom manufacturing is not “one size fits all” and can be simple or complex, such as producing a single component to a final finished product.
Continue reading “Custom Manufacturing: Translating Research into Product”The Largest Maxprep Liquid Handler Installation Ever: Kigali Rwanda, 2022
“It was just a sea of Promega everywhere,” says Rebecca Roberts, a Promega Field Applications Scientist. “Floor to ceiling, piled up with Maxwell instruments, Maxprep Liquid Handlers, all the accessories and consumables…”
In her role on the Field Application Scientists team, Rebecca travels the United States installing the Maxprep Liquid Handler in customer labs and training scientists to operate the system and incorporate it into their workflow. This instrument automates the pre- and post-processing steps in a nucleic acid purification workflow. It’s a large and sophisticated instrument that takes up roughly four feet of lab bench space and weighs up to 220 pounds. It is intended for research use only, but during the COVID-19 pandemic, the Maxprep Liquid Handler, Maxwell RSC 48 Instrument, and several Maxwell purification kits were recommended for nucleic acid extraction protocols in the CDC 2019-Novel Coronavirus Real-Time RT-PCR Diagnostic Panel Emergency Use Authorization (EUA).
When an instrument is sold, Rebecca and a Service Engineer spend three days on-site installing it and training a small group of staff to use it. One Maxprep instrument at a time is typical. On rare occasions, Rebecca might install two on a single trip. However, in 2022, Rebecca joined a multinational team of Promega scientists and engineers in Kigali, Rwanda for an order that was anything but typical.
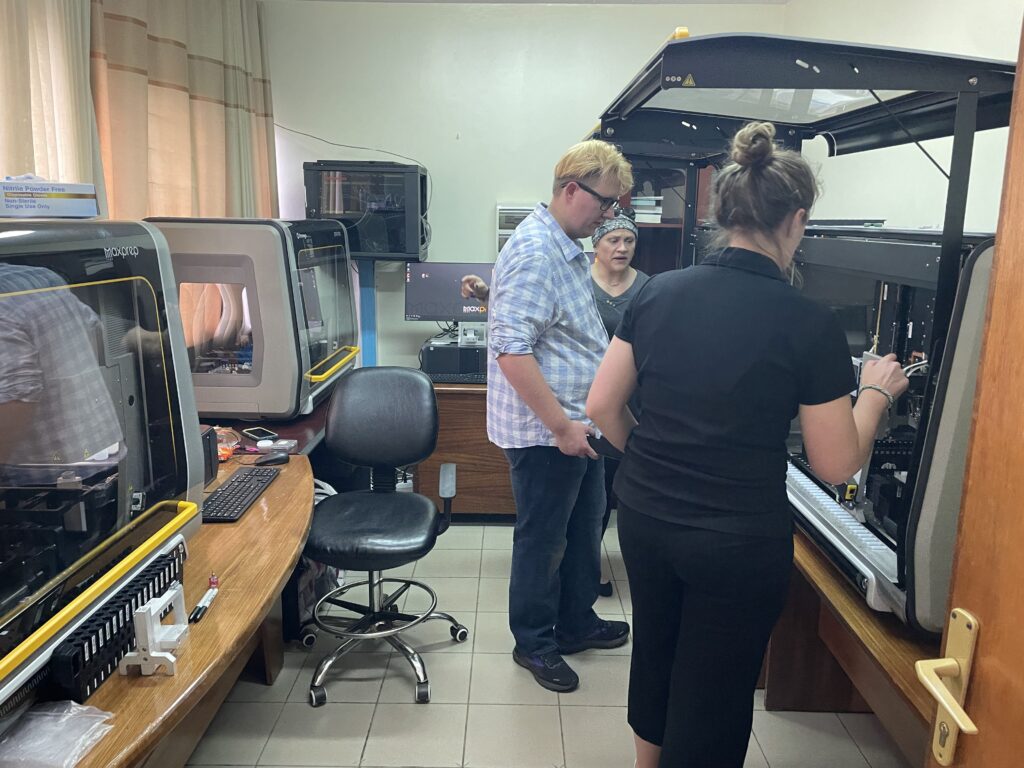

“We knew a large order from this customer was a possibility,” Rebecca says, “But I certainly wasn’t expecting an order of ten.”
This was the largest installation of Maxprep instruments Promega has ever seen from a single order. The customer also had a hard deadline that required delivery, installation and training to be complete in only six weeks – half the time usually quoted for a single instrument.
In the end, ten Maxprep instruments were installed at the National Reference Laboratory in Kigali, and more than twenty people were trained to use the systems for RNA extraction to support COVID-19 testing at a major international meeting. The order was a success, but that six week journey was a wild ride that depended on the hard work and dedication of Promega teams on both sides of the Atlantic.
And the impact of this work is still growing.
Continue reading “The Largest Maxprep Liquid Handler Installation Ever: Kigali Rwanda, 2022”Shifting Gears: Repurposing Instruments for Changing Needs

The thought of an expensive instrument falling out of use and gathering dust on the shelf is enough to bring a tear to any lab manager’s eye. An instrument that once served a key purpose and now functions only as a “paperweight” is a tragic waste of valuable resources. Fortunately, it is sometimes possible to breathe new life into neglected tools and to retrofit or repurpose equipment to meet the new needs that will inevitably arise in a changing lab environment.
Continue reading “Shifting Gears: Repurposing Instruments for Changing Needs”Promega qPCR Grant Funds Genetic Database for Antarctic Krill
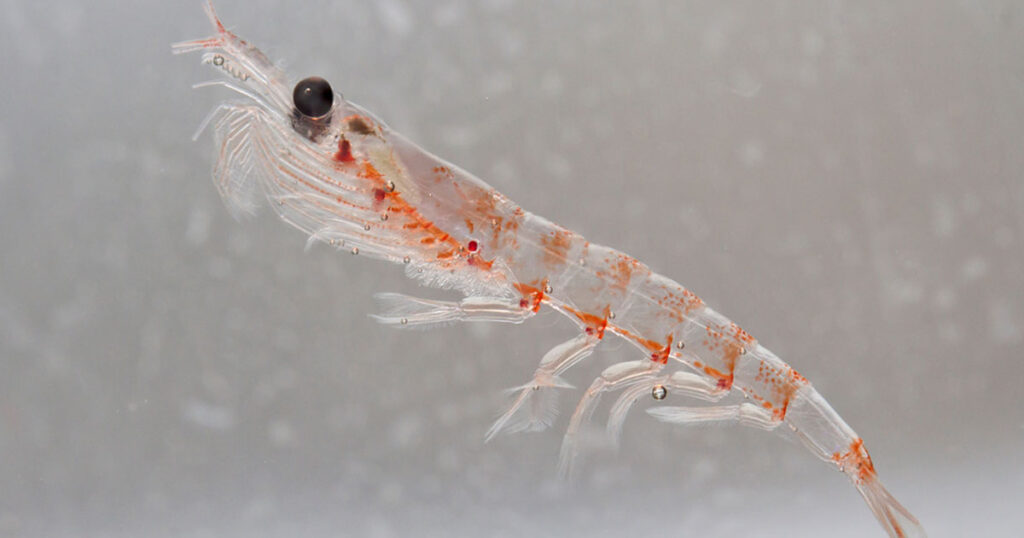

Boasting a biomass of roughly 400 million tons, Antarctic krill are a key source of food for a wide array of marine life, including sea birds, seals, penguins and whales. As with the rest of the oceanic ecosystem, krill are subject to rapidly shifting climate conditions, prompting scientists to seek a deeper understanding of how they might adapt to a changing environment.
Facing a general lack of genetic information on the species, Professors Cristiano De Pittà and Gabriele Sales from the Department of Biology at the University of Padova in Italy set out to define the krill transcriptome, or sequences of ribonucleic acid (RNA), and in doing so facilitated the discovery of key gene sequences that may play important roles in krill reproduction and survival.
In recent years, there have been concerns about potential impacts to the krill population from ocean warming and commercial fishing operations. Mapping the krill transcriptome may offer scientists crucial insight into the effects of climate change and anthropogenic activity on the dynamics of the Antarctic ecosystem. Doing so is no small feat. Though krill may be miniscule, their genome is 15 times the size of the human genome.
To this end, the research groups of De Pittà and Sales established the database KrillDB, providing a single resource where scientists can access a comprehensive catalogue of krill genes and RNA transcripts. This database represented a powerful bioinformatic tool for examining molecular processes in krill. Funded in part by the Promega 2019 qPCR Grant Program, researchers subsequently rolled out an updated database, KrillDB2, which includes improvements to the quality and breadth of the sequences covered and the information associated. Their corresponding study, published summer 2022 in Scientific Reports, identified a series of genes involved in the krill molting cycle, the reproductive process and sexual maturation, and included never-before reported insights into the expression of microRNA precursors and their effect on krill physiology.
The 2019 Promega qPCR Grant Program offered recipients $10,000 in free PCR reagents and related products, as well as access to Promega technical services and training teams.
Researcher and awardee Alberto Biscontin said of the grant’s impact on their project: “RNA sequencing approaches allow us to determine the level of expression of thousands of genes with a single experiment. The standard in the field is to define transcript expression levels by quantitative RT-PCR to technically validate RNA-seq results. We have been relying on the GoTaq® qPCR solutions by Promega for years.” He added, “We have used the GoTaq® 1-Step RT-qPCR System to compare the level of expression of candidate genes with those obtained from RNA-seq analysis. This allowed us to verify at any time the reliability of our bioinformatics pipelines.”
In the future, researchers plan to maintain the KrillDB2 database with the latest genome and transcriptome sequencing data, to provide the most comprehensive integrative analysis possible. They intend to develop a multi-crustaceans database to support future comparative genomics studies. The KrillDB2 database may also serve as a model to develop other databases for similar species.
Learn more about the GoTaq® 1-Step RT-qPCR System.



