“Are you going to the talk?”
The refrain regularly echoes through the halls of every academic lab building. During our education, we’re treated to a non-stop supply of speakers on every subject we can imagine. Prestigious speaker series gave us chances to hear from some of the world’s most prominent experts on subjects that would shape scientific pursuits for the next decade and beyond. When we leave academia, however, it can be difficult to find those same opportunities to learn. Sure, there are lab meetings and conferences, but when can you be treated to a renowned expert giving a talk just down the hall?
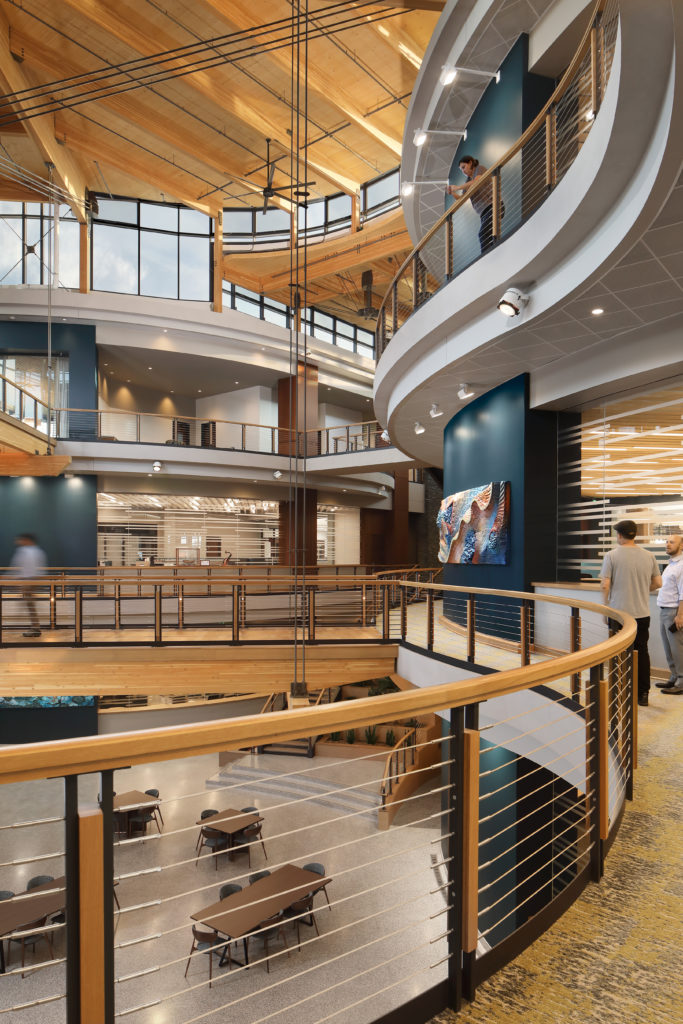
Promega Head of Biology Frank Fan aimed to address that problem when he developed a plan for the Kornberg Innovation Seminars (KIS), a recurring speaker series to be held in the new home for Promega R&D. Kornberg Center is an environment where Promega scientists are challenged to think outside-the-box and anticipate the challenges life science researchers will be facing tomorrow. Frank believed that opportunities to learn from a wide variety of guest experts would be critical for inspiring that type of thinking.
“Promega R&D focuses on understanding scientists’ needs and providing novel solutions,” Frank says. “The KIS program is about helping us achieve that vision.”
Continue reading “Kornberg Innovation Seminars: Inspiring Creativity in Promega R&D”