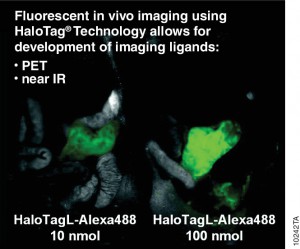
 Antibodies labelled with radioisotopes or the sequential administrationof an antibody and a radioactive secondary agent facilitate the in vivo detection and/or characterisation of cancers by positron emission tomography (PET) or by single-photon emission computed tomography (SPECT) imaging.
Antibodies labelled with radioisotopes or the sequential administrationof an antibody and a radioactive secondary agent facilitate the in vivo detection and/or characterisation of cancers by positron emission tomography (PET) or by single-photon emission computed tomography (SPECT) imaging.
There are drawbacks to both methods, including prolonged exposure to radiation and ensuring that both the antibody and the radiolabelled secondary agent are suitably designed so that they bind rapidly upon contact at the tumor.
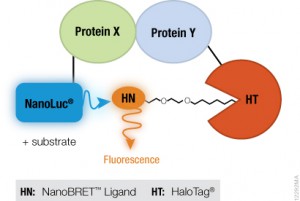
A recent publication (1) investigated a alternative method utilizing the HaloTag® dehalogenase enzyme HaloTag® is a dehalogenase enzyme (33 kDa) that contains an engineered cavity designed to accommodate the reactive chloroalkane group of a HaloTag® ligand (HTL). Upon entering the enzyme cavity, the terminal chlorine atom rapidly undergoes nucleophilic displacement, and a covalent adduct is formed, effectively anchoring the HaloTag® ligand in a precise location.
Three new HaloTag® ligands were synthesized and each labelled with the SPECT radionuclide indium-111 111In-HTL-1 and the dual-modality HaloTag® ligands,111In-HTL-2 and111;In-HTL-3 containing TMR which allows complementary imaging data).
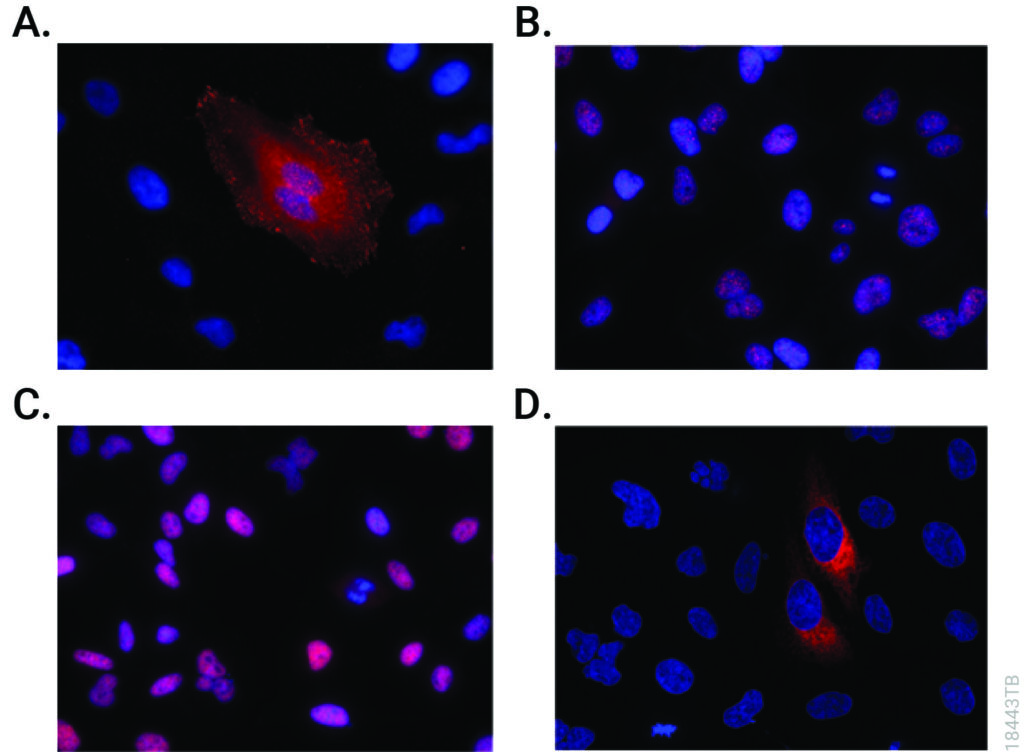
For the validation of the pretargeting strategy based on these HaloTag® ligands, the target human epidermal growth factor receptor 2 (HER2)was selected. Trastuzumab (Herceptin®) was selected as the primary targeting agent and was modified with HaloTag® protein via the trans-cyclooctene/tetrazine ligation.
All three 111In-labelled HaloTa®g ligands exhibited significantly higher binding to the HER2 expressing when compared to negative controls.
Literature Cited
Knight, J. C et al.(2015) Development of an enzymatic pretargeting strategy for dual-modality imaging. Chem. Commun. 51, 4055–8.

 Designing and implementing a new assay can be a challenging process with many unexpected troubleshooting steps. We wanted to know what major snags a scientist new to the
Designing and implementing a new assay can be a challenging process with many unexpected troubleshooting steps. We wanted to know what major snags a scientist new to the  Antibodies labeled with small molecules such as fluorophore, biotin or drugs play a critical role in various areas of biological research,drug discovery and diagnostics. There are several limitations to current methods for labeling antibodies including the need for purified antibodies at high concentrations and multiple buffer exchange steps.
Antibodies labeled with small molecules such as fluorophore, biotin or drugs play a critical role in various areas of biological research,drug discovery and diagnostics. There are several limitations to current methods for labeling antibodies including the need for purified antibodies at high concentrations and multiple buffer exchange steps.
 Antibodies labelled with radioisotopes or the sequential administrationof an antibody and a radioactive secondary agent facilitate the in vivo detection and/or characterisation of cancers by positron emission tomography (PET) or by single-photon emission computed tomography (SPECT) imaging.
Antibodies labelled with radioisotopes or the sequential administrationof an antibody and a radioactive secondary agent facilitate the in vivo detection and/or characterisation of cancers by positron emission tomography (PET) or by single-photon emission computed tomography (SPECT) imaging.