Tumor cells are characterized by many features: including uncontrolled proliferation, to loss of contact inhibition, acquired chromosomal instability and gene copy number changes among them. Some of those copy number changes are site-specific, but very little is known about the mechanisms or proteins involved in creating site-specific copy number changes. In a recently published Cell paper, Black and colleagues, propose a mechanism for site-specific copy number variations involving histone methylation proteins and replication complexes.
Previous work from Klang et al. had shown that local amplification of chromosomal regions occurs during S phase and that chromatin structure plays a critical role in this amplification (2), and other work by Black and colleagues (3) implicated KDM4A in changing timing of replication by altering chromatin accessibility in specific regions. Other research also had shown that KDM4A protein levels influence replication initiation and that KDM4A has a role in some DNA damage response pathways (4,5). Looking at the body of work, Black et al. hypothesized that KDM4A, with its roles in replication, might possibly provide link into the mechanism of site-specific copy number variation in cancer.
Is KDM4A copy number altered in tumor cells?
The first question that Black and colleagues asked was: Is KDM4A over expressed in tumor cells? To address this question they looked at samples from The Cancer Genome Atlas (TCGA). They assayed 1,770 primary tumor samples from 8 cancer types and found that in 18.9% of them KDM4A gene copy number was increased, and in 22.1% of them, KDM4A gene copy number was decreased. The changes in copy number were accompanied by the predicted changes in protein expression. When they looked at ovarian cancer samples specifically, they found that 46% of ovarian tumor samples showed KDM4A amplification and that amplification in ovarian cancer was associated with a shortened median time to death.
Does KDM4A over expression promote genomic instability?
They next asked what the phenotypic effect was in cells in which KDM4A was over expressed. They stably over expressed KDM4A in genomically stable (as indicated by karyotype), immortal but not transformed, RPE-hTERT cells (human retinal pigmented epithelium cells immortalized with human telomerase (6). The stably transfected cells expressed KDM4A at 2–3 times above levels observed in control cells; this level of over expression did not lead to global genomic instability. However, Black et al. were able to detect increased copy number of 1q12h in 14% of the stably transformed cells. This was not a result of increased copies of the entire long arm (1q) of chromosome 1, because the telomere for that arm did not show increased copy number.
Is catalytically active KDM4A required for 1q12h copy gain?
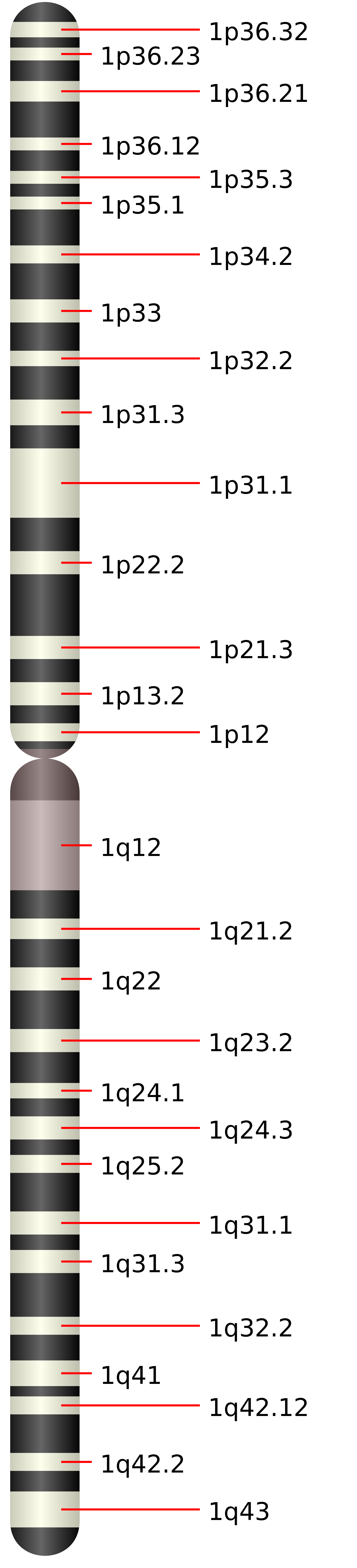
After determining that over expression of KDM4A could lead to site-specific copy number changes, Black et al. next sought to determine what parts of the KDM4A protein were necessary for this effect. They over expressed catalytically active and catalytically inactive forms of KDM4A. Their experiments showed that transient over expression of catalytically active KDM4A resulted in 1q12h copy number gains, while transient over expression of catalytically inactive KDM4A did not. Furthermore, this copy number gain occurred within 24 hours and was specific to KDM4A: it was not seen with histone lysine demethylases KDM4B, KDM4C or KDM4D, the family members most closely reated to KDM4A.
Black et al. next expressed histones with methionine replacing specific lysine residues to see if these altered histones would promote 1q12h copy number gain. Expression of histone 3 (H3) with methionine substituted for either lysine at position 9 or 3 did promote 1q12h copy number gain in a manner similar to that observed with KDM4A over expression. This copy number gain could be suppressed by the overexpression of the Suv39h1 methyltransferase.
HP1g (heterochromatin protein 1) is a chromatin binding protein that is known to antagonize KDM4A changes in the cell cycle. HP1g binds to methylated H3K9, and blocks access to KDM4A and other DNA replication proteins. KDM4A demethylates H3K9, allowing replication proteins access to DNA, and progression of S phase. In experiments in which HP1g and KDM4A were cotransfected and over expressed simultaneously, HP1g could antagonize KDM4A-induced 1q12h site-specific copy number gain. However, if the HP1g was added after the KDM4A had been previously over expressed, the site-specific copy number gain was not blocked, suggesting that once an altered chromatin structure was established HP1g was not sufficient to block the copy number changes.
Is the 1q12h copy number change stably inherited?
To determine the pattern of inheritance of the 1q12h copy number gain, Black et al. isolated single-cell clones from the RPE cell line that stably over expressed KDM4A. They performed FISH analysis on each of the clones to see if they had the copy number gain. Stable inheritance would predict 100% of the clones positive for the copy number gain. However, what they observed was that 1.5–37% of the clones were positive for the 1q12h copy number gain, indicating that it was not stably inherited.
When Black et al. looked at the cells during various phases of the cell cycle. Using various cell cycle arrest agents, they were able to determine that the copy number gain of 1q12h occurred during S phase and was eliminated by late G2 phase. This means that the amplified DNA did not become incorporated into the genome and was only transiently present in the cell.
What are the molecular mechanisms underlying KDM4A-induced site-specific copy number variation?
To understand more about the molecular players in site-specific copy number variation, Black et al. used a HaloTag-KDM4A fusion to identify KDM4A interacting proteins. The proteins identified were involved in replication, particularly rereplication and included proteins of the minichromosome maintenance complex (MCM 2, 3 and 7) and DNA polymerase subunits. The isolation of these players suggests that KDM4A promotes rereplication in 1q12.
To investigate the re-replication hypothesis, Black et al. performed cesium chloride density gradient centrifugation of DNA. They labeled cells for less than one complete cell cycle, enriching for heavy-light (replicated DNA). They did not detect a peak of heavy-heavy DNA (H:H); however, when they looked at pooled the H:H DNA fractions and assayed for specific regions, they found a 7-fold enrichment for targets in 1q12 region in KDM4A-overexpressing cells.
The Model and the Questions
From these results it is clear that transient KDM4A overexpression can cause site-specific copy number variation during a single cell cycle, presumably by leading to an “open” chromatin structure that leads to rereplication by recruiting DNA polymerase subunits and other replication-associated proteins. If the sites of rereplication contain oncogenes, like 1q12 does, the “extra” copies of these genes could promote tumorigenesis.
Many questions remain. For instance, in their initial investigation of the TCGA, the authors note not only increased KDM4A protein but also amplification of KDM4A. The initial amplification event for KDM4A is not understood. Additionally, we do not understand how the rereplicated regions are removed during the exit from S phase—what is the pathway for degradation of this extrachromosomal DNA?
What is certain is that Black et al. have opened an exciting new arena for cancer research with new avenues to pursue for therapeutic targets and better understanding of intractable cancers. It will be exciting to see what research in this area yields over the next few years.
References
- Black J., Manning A., Van Rechem C., Kim J., Ladd B., Cho J., Pineda C., Murphy N., Daniels D. & Montagna C. & (2013). KDM4A Lysine Demethylase Induces Site-Specific Copy Gain and Rereplication of Regions Amplified in Tumors, Cell, 154 (3) 541-555. DOI: 10.1016/j.cell.2013.06.051
- Klang, L. et al. (2010) Specific replication origins promote DNA amplification in fission yeast. J. Cell Sci. 123, 3047–51.
- Black, J.C. et al. (2010) Conserved antagonism between JMJD2A/KDM4A and HP1g during cell cycle progression. Mol. Cell 40, 736–48.
- Van Rechem, C. et al. (2011) The SKP1-Cul1-F-box and leucine-rich repeat protein 4 (SCF-FbxL4) ubiquitin ligase regulates lysine demethylase 4A (KDM4A)/Jumonji domain-containing 2A (JMJD2A) protein. J. Biol. Chem. 286, 30462–70.
- Mallet, F.A. et al. (2012) RNF-8 and RNF-168-dependent degradation of KDM4A/JMJD2A triggers 53BP1 recruitment to DNA damage sites. EMBO J. 31, 1865–78.
- Jiang, X.R. (1999) Telomerase expression in human somatic cells does not induce changes associated with a transformed phenotype. Nat. Gen. 21, 111–4.
Michele Arduengo, PhD
Latest posts by Michele Arduengo, PhD (see all)
- Why BRETSA™ Target Engagement Matters for Drug Discovery - March 10, 2026
- What Shelter Dogs Can Tell Us About Emerging Zoonotic Diseases - October 2, 2025
- Developing an Experimental Model System to Understand the Tumor Microenvironment of Melanoma Brain Metastases - September 4, 2025
