The pregnane X receptor (PXR) is a nuclear receptor known to regulate expression of cytochrome P450 (CYP450) drug-metabolizing enzymes (1). PXR has even been designated the “master xenosensor” due to its ability to upregulate cellular levels of a variety of drug-metabolizing enzymes in response to drugs and foreign chemicals. Elevated levels of CYP450 enzymes can elicit alterations in the pharmacokinetics of co-administered drugs, which can result in adverse drug-drug interactions (DDI) or diminished bioavailability. By assessing PXR activation and CYP450 enzyme induction early in the drug development process, many companies hope to reduce late-stage clinical failures and minimize the high costs associated with bringing a new drug to market.
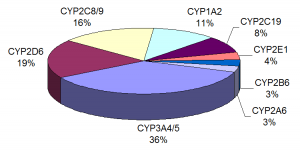
A paper by Shukla et al. (2) examined over 2,800 clinically used drugs for their ability to activate human PXR (hPXR) and rat PXR (rPXR), induce human cytochrome P450 3A4 enzyme (CYP3A4) at the cellular level, and bind hPXR at the protein level. Several studies have identified PXR as playing a key role in regulating the expression of CYP3A4, an enzyme involved in the metabolism of more than 50% of all drugs prescribed in humans. Since PXR activation and CYP3A4 induction have an impact on drug metabolism and pharmacokinetics, the authors wanted to obtain data that would be valuable in understanding structure-activity relationships (SARs), the connection between chemical structure and biological activity, when prioritizing new molecular entities (NMEs) for further in vitro and in vivo studies.
The authors used quantitative high-throughput screening (qHTS) to profile clinical drugs using cell-based assays for hPXR, rPXR and human CYP3A4 activation. They also performed PXR ligand-binding domain (LBD) assays on these same drugs. Nuclear receptor activation, CYP3A4 –mediated substrate metabolism and cellular toxicity were measured in whole-cell systems developed by Puracyp (Carlsbad, CA). Essentially, a stable cell line, designated DPX2™, was derived from HepG2 cells cotransfected with an expression vector containing the full-length coding region of hPXR, a PXR response element (PXRE)-luciferase gene and a CYP3A4 enhancer element upstream from the luciferase reporter gene. The PXRE is required for binding PXR in regulatory regions of target genes before their activation. A similar stable transfectant was created for rPXR using a rodent hepatoma cell line. Both cell lines were used in the study as a means to uncover species-specific differences when trying to extrapolate animal PXR activation data to humans.
The three transactivation reporter gene assays and the PXR LBD assay were performed in separate 1536-well, white opaque plates.
- To measure PXR activation in response to varying drug concentrations in the DPX2™ and rPXR cell lines, the luciferase reporter gene activity (3) was determined using the Promega ONE-Glo™ Luciferase Assay.
- CYP3A4 metabolic activity was assessed in both cell lines after drug treatment using the Promega P450-Glo™ CYP3A4 Assay with Luciferin-PPXE, a luminogenic CYP3A4-specific substrate. Upon substrate cleavage, luciferin is released, producing light proportional to enzyme activity in a luciferase reaction.
- The hPXR LBD assay employed competitive Time-Resolved Fluorescence Resonance Energy Transfer (TR-FRET) with a fluorescein-labeled PXR ligand.
- Cell viability was measured in separate plates with a luciferase-coupled ATP assay to help normalize data to the number of metabolically active cells.
A subset of compounds was retested to confirm sample integrity and assay reproducibility. In addition, the authors looked at CYP3A4 enzyme induction in cryopreserved, human primary hepatocytes after drug treatment using the Promega P450-Glo™ CYP3A4 Assay containing the highly sensitive Luciferin-IPA substrate. Hepatocyte cell viability (ATP quantitation) was assessed in the same wells after a wash step using the CellTiter-Glo® Luminescent Cell Viability Assay from Promega. Data obtained in DPX2™ cells and human hepatocytes showed a good correlation among therapeutics known to induce CYP3A4 enzyme, further validating the usefulness of the DPX2™ cell line to predict DDI.
Shukla et al. were able to identify a few common chemical scaffolds from their SAR analysis by categorizing compounds into eight activity clusters defined by the results observed across the four different assays. For example, dihydropyridine calcium channel blockers showed activity in all four assays, while steroids and fatty acids demonstrated activity in only one out of the four assays (see paper for details). Although there were significant species differences in potency, primary screening of known drugs for PXR and CYP3A4 induction, followed by SAR analysis, provided data that could potentially eliminate undesirable DDI chemotypes early in the drug discovery process or during rational drug design.
By simply supplying the test compounds, researchers can complete a DDI screen in 2 days with a total of 3–6 hours of hands-on time. Similar multiplex kits for assessing activation of human Aryl hydrocarbon receptor (AhR) are also available from Puracyp.
References
- Chu, V., et al. (2009) In vitro and in vivo induction of cytochrome P450: A survey of the current practices and recommendations: A Pharmaceutical Research and Manufacturers of America perspective. Drug Metab. Dispos. 37, 1339–54.
- Shukla, S.J. et al. (2011) Identification of clinically used drugs that activate pregnane X receptors. Drug Metab. Dispos. 39, 151–9.
- Raucy, J.L. and Lasker, J.M. (2010) Current in vitro high throughput screening approaches to assess nuclear receptor activation. Curr. Drug Metab. 11, 806–14.

Congratulations! Promega has been nominated in several categories for the 2011 Life Science Awards! See you next month in Washington, DC! You can see the full list of nominees at http://www.lifescienceindustryawards.com.