Protein-DNA interactions are fundamental processes in gene regulation in a living cells. These interactions affect a wide variety of cellular processes including DNA replication, repair, and recombination. In vivo methods such as chromatin immunoprecipitation (1) and in vitro electrophoretic mobility shift assays (2) have been used for several years in the characterization of protein-DNA interactions. However, these methods lack the throughput required for answering genome-wide questions and do not measure absolute binding affinities. To address these issues a recent publication (3) presented a high-throughput micro fluidic platform for Quantitative Protein Interaction with DNA (QPID). QPID is an microfluidic-based assay that cam perform up to 4096 parallel measurements on a single device.
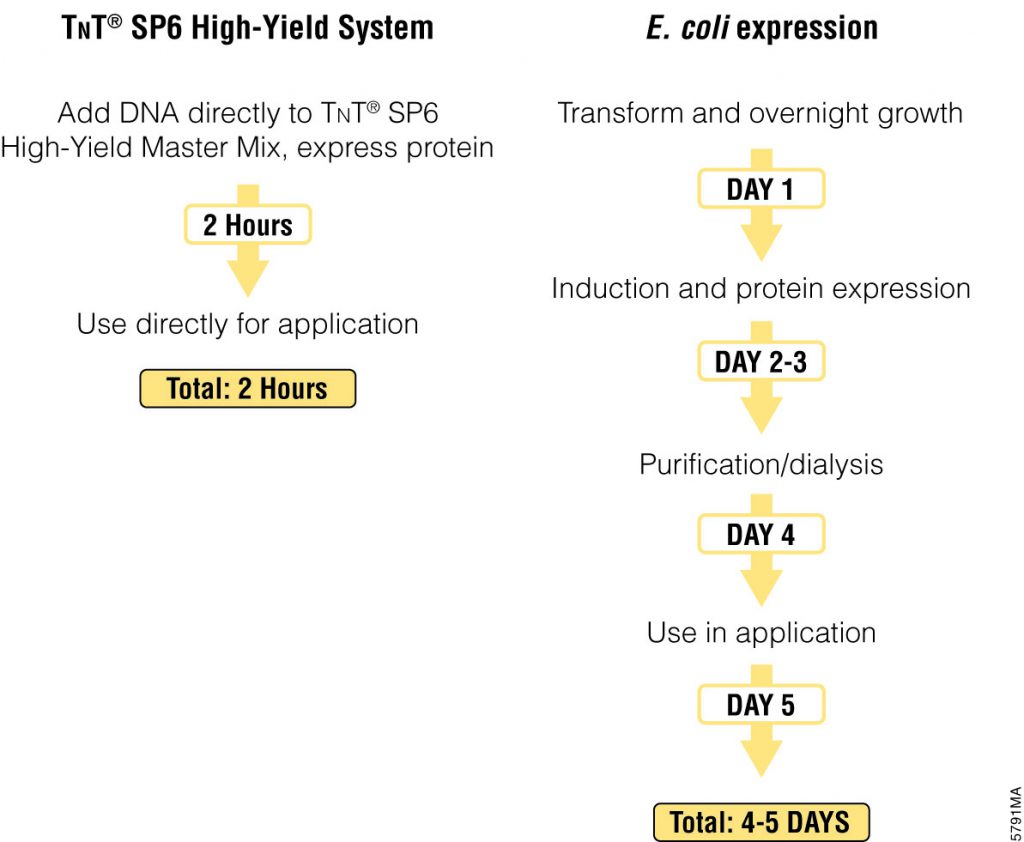
The basic elements of each experiment includes oligonucleotides that were synthesized and hybridized to a Cy5-labeled primer and extended using Klenow. All transcription factors that were evaluated contained a 3’HIS and 5’ cMyc tag and were expressed in rabbit reticulocyte coupled transcription and translation reaction (TNT® Coupled Reticulocyte Lysate). Expressed proteins are loaded onto to the QIPD device and immobilized. In the DNA binding assay the fluorescent DNA oligonucleotides are incubated with the immobilized transcription factors and fluorescent images taken. To validate this concept the binding of four different transcription factor complexes to 32 oligonucleotides at 32 different concentrations was characterized in a single experiment. In a second application, the binding of ATF1 and ATF3 to 128 different DNA sequences at different concentrations were analyzed on a single device.
Literature Cited
- Ren, B. et al. (2007) Genome-wide mapping of in vivo protein-DNA binding proteins. Science 316, 1497–502.
- Garner, M.M. (1981) A gel electrophoresis method for quantifying the binding of proteins to specific DNA regions. Nuc. Acids. Res. 9, 3047-60.
- Glick,Y et al. (2016) Integrated microfluidic approach for quantitative high throughput measurements of transcription factor binding affinities. Nuc. Acid Res. 44, e51.


![Bubonic plague victims in a mass grave in 18th century France. By S. Tzortzis [Public domain], via Wikimedia Commons](https://upload.wikimedia.org/wikipedia/commons/1/1b/Bubonic_plague_victims-mass_grave_in_Martigues%2C_France_1720-1721.jpg)