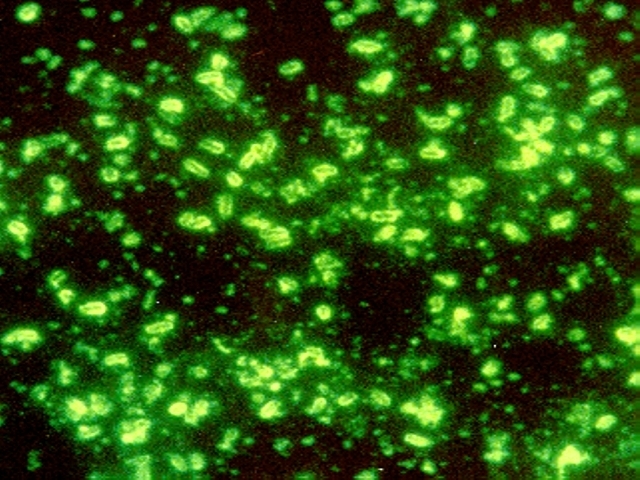
Following the discovery of Mimivirus (1) the first virus with a particles large enough to be visible under the light microscope, two additional “giant” viruses infecting Acanthamoeba have been discovered Pandoravirus (2) and Pithovirus sibericum (3), the latter from a 30,000 year old Siberian permafrost. A fourth type was recently isolated from the same sample of permafrost by Legendre et al, and named Mollivirus sibericum (4).
Mollivirus sibericum has an approximately spherical virion (0.6 µm diameter) with a 651kb GC-rich genome that encodes 523 proteins. To further characterize the virus the researchers performed transcromic- and proteomic-based time course experiments.
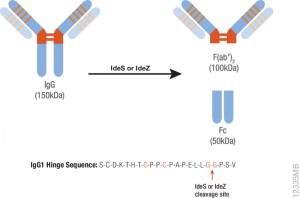
For the particle proteome and infectious cycle analysis, proteins were extracted and then run a 4–12% polyacrylamide gel, and trypsin digests were performed in-gel before nano LC-MS/MS analysis of the resulting peptides. Proteomic studies of the particle showed that it lacked an embarked transcription apparatus, but revealed an unusual presence of many ribosomal and ribosome-related proteins.
When the researchers explored the proteome during the course of an entire infectious cycle, the relative proportions of Mollivirus-, mitochondrion-, and Acanthamoeba encoded proteins were found to vary consistently with an infectious pattern that preserved the cellular host integrity as long as possible and with the release of newly formed virus particles through exocytosis.
In an interesting footnote, the authors of this study point out the fact that two different viruses retain their infectivity in prehistorical permafrost layers should be a concern in the context of global warming and the potential to expose humans to primeval viruses.
References
1. La Scola, B. et al. (2003) A giant virus in amoebae. Science 299, 2033.
2. Philippe, N. et al. (2013) Pandoraviruses. Amoeba virus with genomes up to 2.5Mb reaching that of parasitic eukaryotes. Science 341,281–6.
3. Legendre, M. et al. (2014) Thirty thousand year old distant relative of giant icosahedral DNA viruses with a pandoravirus morphology. Proc.Natl. Acad. Sci. 111, 4274–9.
4. Legendre, M. et al. (2015) In depth study of Mollivirus sibercum, a new 30,000 year old giant virus infecting Acanthamoeba. Proc. Natl. Acad. Sci. 112, E5327–35 (online).