In today’s post, guest blogger, Martha O’Brien, PhD, provides a preview of her upcoming AAI poster and block symposium talk on the inflammasome, caspase-1 activity and pyroptosis.
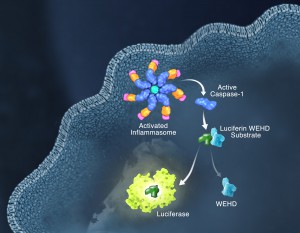
Responding rapidly to microbial pathogens and damage-associated molecular markers is critical to our innate immune system. Caspase-1 is pivotal in this process leading to processing and release of essential cytokines and an immunogenic form of cell death, termed pyroptosis. Upon sensing pathogen-associated and damage-associated molecular patterns (PAMPs and DAMPs), innate immune cells form inflammasome protein complexes that recruit and activate caspase-1 (canonical inflammasomes). In addition, other inflammatory caspases, 4 and 5 in humans and 11 in mice, directly bind bacterial lipopolysaccharides (LPS), triggering pyroptosis (non-canonical inflammasome). LPS-triggered non-canonical inflammasomes in mice and humans ultimately lead to canonical inflammasome engagement and caspase-1 activation (1–3). Caspase-1 was originally termed interleukin converting enzyme (ICE) for its well-established role in processing IL-1ß and IL-18, two important inflammation cytokines. How caspase-1 mediates pyroptosis is less well understood, but is beginning to be delineated. Recently, a substrate of the inflammatory caspases, gasdermin D, was identified and its processed fragment, gasdermin-N domain, was shown to be required for pyroptosis in non-canonical inflammasome circumstances (4, 5). The precise role of gasdermin D in canonical inflammasome-triggered pyroptosis is still under investigation. Linking inflammatory caspases directly to pyroptosis is a notable step in understanding the mechanism of this important form of cell death.
Pyroptosis is clearly one means of releasing processed IL-1ß and IL-18 from the cell. However depending on the cell type and stimulus, there is evidence for inflammasome engagement, caspase-1 activation, and release of IL-1ß in the absence of cell death (6, 7). On the flip-side there is also evidence for caspase-1 mediated pyroptosis that helps clear bacteria, independent of IL-1ß and IL-18 involvement (8). To enable further studies on the inflammasome and in particular, assessing the connections between caspase-1 activation, pyroptosis, and cytokine release, Promega developed a new tool to conveniently monitor caspase-1 activation, the Caspase-Glo® 1 Inflammasome Assay. This bioluminescent, plate-based assay is used to measure caspase-1 activity directly in cell cultures or to monitor released caspase-1 activity in culture medium from treated cells. This flexibility allows easy multiplexing to monitor all three outcomes of inflammasome stimulation; caspase-1 activity, pyroptosis, and release of IL-1ß and IL-18. Caspase-1 activation typically is monitored indirectly with western blots of processed caspase-1. Now the activity of the enzyme can be monitored directly, providing accurate information on temporal aspects of the inflammasome. The assay can be readily combined with real-time measures of cell death (e.g., CellTox™ Green Cytotoxicity Assay) and some of the culture medium can be removed for IL-1ß/IL-18 assessment, leaving the cells and remaining culture medium for caspase-1 activity measurements.
References
- Schmid-Burgk et al. (2015) Caspase-4 mediates non-canonical activation of the NLRP3 inflammasome in human myeloid cells. J. Immunol. 45, 2911–7.
- Baker et al. (2015) NLRP3 inflammasome activation downstream of cytoplasmic LPS recognition by both caspase-4 and caspase-5. J. Immunol. 45, 2918–26.
- Ruhl, S. and P. Broz (2015) Caspase-11 activates a canonical NLRP3 inflammasome by promoting K+ Eur. J. Immunol. 45, 2927–36.
- Shi et al. (2015) Cleavage of GSDMD by inflammatory caspases determines pyroptotic cell death. Nature 526, 660–5.
- Kayagaki et al. (2015) Caspase-11 cleaves gasdermin D for non-canonical inflammasome signaling. Nature 526, 666–71.
- Gaidt et al. (2016) Human monocytes engage an alternative inflammasome pathway. Immunity 44, 833–46.
- Chen et al. (2014) The neutrophil NLRC4 inflammasome selectively promotes IL-1ß maturation without pyroptosis during acute. Salmonella Cell Reports 8, 570–82.
- Miao et al. (2010) Caspase-1-induced pyroptosis is an innate immune effector mechanism against intracellular bacteria. Nature Immunology 11, 1136–42.

 Recruiters aren’t pessimists, but throughout the years we have become more cautious and maybe a little suspicious. Many of us interviewed enough candidates that we have come to approach each new person with a “trust but verify” mentality. I’m very trusting in my personal life, but at work, my job is to be a detective. I follow clues to dig up the good, the bad, and the ugly.
Recruiters aren’t pessimists, but throughout the years we have become more cautious and maybe a little suspicious. Many of us interviewed enough candidates that we have come to approach each new person with a “trust but verify” mentality. I’m very trusting in my personal life, but at work, my job is to be a detective. I follow clues to dig up the good, the bad, and the ugly.

![Fig 4. Four point MMOA screen for tideglusib and GW8510. Time dependent inhibition was evaluated by preincubation of TbGSK3β with 60 nM tideglusib and 6 nM GW-8510 with 10μM and 100μM ATP. (A). Tideglusib [60 nM] in 10μM ATP. (B). GW8510 [60 nM] in 10μM ATP. (C.) Tideglusib [60 nM] at 100μM ATP. (D.) GW8510 [60 nM] at 100μM ATP. All reactions preincubated or not preincubated with TbGSK3β for 30 min at room temperature. Experiments run with 10μM GSM peptide, 10μM ATP, and buffer. Minute preincubation (30 min) was preincubated with inhibitor, TbGSK3β, GSM peptide, and buffer. ATP was mixed to initiate reaction. No preincubation contained inhibitor, GSM peptide, ATP, and buffer. The reaction was initiated with TbGSK3β. Reactions were run at room temperature for 5 min and stopped at 80°C. ADP formed was measured by ADP-Glo kit. Values are mean +/- standard error. N = 3 for each experiment and experiments were run in duplicates. Control reactions contained DMSO and background was determined using a zero time incubation and subtracted from all reactions. Black = 30 min preincubation Grey = No preincubation.](https://www.promegaconnections.com/wp-content/uploads/2016/04/journal.pntd_.0004506.g004-243x300.png)

 When Dave Romanin came to work for Promega he was fresh out of school with a degree in bacteriology. His plan was to work for a year in manufacturing and then go back to graduate school. But in the end, he didn’t go. There was no incentive, he explains, for him to spend five years in graduate school making little to no money. He didn’t want to write grants or run his own lab, and he enjoyed what he was doing.
When Dave Romanin came to work for Promega he was fresh out of school with a degree in bacteriology. His plan was to work for a year in manufacturing and then go back to graduate school. But in the end, he didn’t go. There was no incentive, he explains, for him to spend five years in graduate school making little to no money. He didn’t want to write grants or run his own lab, and he enjoyed what he was doing.