The understudied kinome represents a major challenge as well as an exciting opportunity in drug discovery. A team of researchers lead by Nathanael Gray at the Dana Farber Cancer Institute was able to partially elucidate the function of an understudied kinase, Doublecortin-like kinase 1 (DCLK1), in pancreatic ductal adenocarcinoma cells (PDAC). The characterization of DCLK1 in PDAC was realized by developing a highly specific chemical probe (1). Promega NanoBRET™ Target Engagement (TE) technology enabled intracellular characterization of this chemical probe.
The Dark Kinome
Comprised of over 500 proteins, the human kinome is among the broadest class of enzymes in humans and is rife with targets for small molecule therapeutics. Indeed, to date, over 50 small molecule kinase inhibitors have achieved FDA approval for use in treating cancer and inflammatory diseases, with nearly 200 kinase inhibitors in various stages of clinical evaluation (2). Moreover, broad genomic screening efforts have implicated the involvement of a large fraction of kinases in human pathologies (3). Despite such advancements, our knowledge of the kinome is limited to only a fraction of its family members (3,4). For example, currently less than 20% of human kinases are being targeted with drugs in clinical trials. Moreover, only a subset of kinases historically has garnered substantial citations in academic research journals (4). As a result, a large proportion of the human kinome lacks functional annotation; as such, these understudied or “dark” kinases remain elusive to therapeutic intervention (4).
If such a large proportion of kinases are deemed integral to cellular function and human disease, why is our knowledge and focus directed to only a small fraction of the family? The answer is complex but often results from a lack of research tools to probe kinase function in a pathophysiological setting. Even for understudied kinases identified as cancer drivers from broad-genome screens, functional annotation and deeper studies on kinase function represent a noteworthy challenge. Since these dark kinases lack functional annotation and available assay tools, highly studied (i.e., “druggable”) kinases are often prioritized over the lesser studied family members.
The Role of Chemical Probes
Chemical probes represent a tool that can help achieve a better understanding of the dark kinome. A chemical probe is a small molecule designed to selectively engage and modulate a specific protein’s function in cells (5). As a result, chemical probes are critical components in research, as they help determine the target protein’s role in complex cellular systems. Kinase chemical probes can be enormously useful tools to illuminate the role of dark kinases in a pathophysiological setting. However, developing selective kinase chemical probes is no trivial endeavor, owing to the highly conserved structure of the nucleotide co-substrate (ATP) binding pocket within the kinase family. Furthermore, the high concentration of intracellular ATP (exceeding Km by orders of magnitude) implies that such probes will generally require strong kinase affinity to achieve intracellular target engagement with commensurate inhibition of kinase activity (6).
Targeting a Dark Kinase Implicated in Cancer
Doublecortin-like kinase 1 (DCLK1) is an example of a dark kinase, where overexpression has been implicated in a variety of cancers, including pancreatic ductal adenocarcinoma (PDAC) (7). Unfortunately, PDAC patient survival rates have not improved in recent years, and our fundamental knowledge of PDAC pathophysiology remains lacking. To query the role of DCLK1 in PDAC proliferation and gene expression, researchers previously utilized nonspecific DCLK1 inhibitors. Such efforts failed to elucidate the specific role of DCLK1 in this context, as collateral inhibition of unrelated kinases confounded interpretations of these experiments (1). This outcome highlights the need for high-quality, selective chemical probes to evaluate the specific effects of DCLK1 inhibition on PDAC cell physiology.
A recent publication by Ferguson et al. (1) at the Dana Farber Research Institute has shed light on the role of DCLK1 inhibition by developing a selective and potent chemical probe for DCLK1. Guided by chemoproteomics and X-ray crystal structures, these researchers were able to iteratively optimize a novel inhibitor that maintained strong inhibitory potency for the primary target DCLK1, while minimizing engagement to collateral kinases. The final optimized inhibitor, DCLK1-IN-1, was highly selective and potent against DCLK1 in cell-free profiling studies. However, demonstrating inhibition in a cell-free setting is only part of what is necessary to develop a chemical probe. To be useful as a chemical probe, DCLK1-IN-1 must be capable of permeating live cells, capable of engaging DCLK1 with high affinity in cells (in the presence of physiological ATP), and be selective for DCLK1. To verify live-cell target engagement under these stringent conditions, the authors developed a novel NanoBRET™ Target Engagement Assay using a population of live cells that expressed a DCLK1-NanoLuc® fusion protein. This expression results in bright, blue light proximal to DCLK1 in the cells. To quantify target engagement, a NanoBRET™ tracer was generated that forms an energy transfer complex with DCLK1-NanoLuc in live cells. When DCLK1-IN-1 engages its target in live cells, the NanoBRET™ tracer is displaced, leading to a loss of energy transfer.
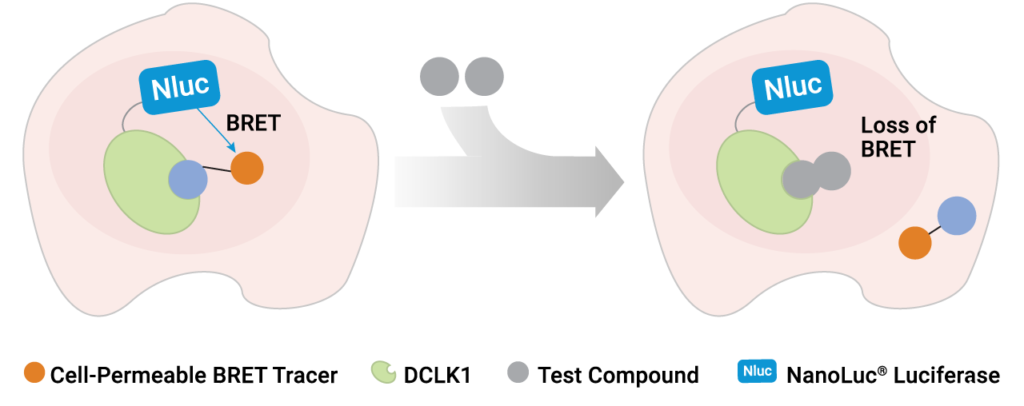
Using a variety of complementary assay methods, such as chemoproteomics, cell-free kinase binding assays and NanoBRET™ target engagement, the authors proved that this novel inhibitor was highly selective for DCLK1 and could engage its intended target in the challenging environment of a live cell. Together, these data showed that DCLK1-IN-1 was not only an inhibitor of DCLK1 but also a suitable chemical probe to explore the role of DCLK-1 activity in cells.
How can DCLK1-IN-1 be used to characterize the pathophysiology of PDAC and does the inhibition of DCLK1 translate into phenotypic effects of PDAC cells or tissues? To answer these questions, the researchers treated PDAC cell lines, as well as patient-derived organoids, with DCLK1-IN-1 and evaluated a number of physiological effects. First, the authors monitored the effects of DCLK1 inhibition on cell proliferation. Interestingly, DCLK1-IN-1 had little effect on PDAC cells in culture, but it had a pronounced antiproliferative effect on patient-derived organoids. This result may imply that 2D cell culture systems are not fully relevant in the context of PDAC pathophysiology. Next, the authors globally evaluated transcriptome and phosphoproteome signatures, and identified a potential role of DCLK1 in cell motility in PDAC.
Conclusions
This research represents a noteworthy example of the power of kinase chemical probes. Here, the team at Dana Farber was able to partially elucidate the functions of an understudied kinase and its vulnerability to chemical inhibition in PDAC. Historically, such outcomes would require knowledge about specific enzymatic substrates and functional characterization. The characterization of DCLK1 in PDAC was realized in the absence of a bona fide kinase substrate, underscoring the value of kinase chemical probes in understudied kinase research. Enabled by a powerful set of new target engagement technologies and the emergence of novel chemical probes, the future is bright for the dark kinome. As a result, we are one step closer to a fundamental understanding of the pathophysiology of a challenging form of pancreatic cancer.
The NanoBRET™ TE Kinase Assays measure the intracellular affinity of test compounds using a multi-well plate format. Learn More
References
- Ferguson, F. M. et al. (2020) Discovery of a selective inhibitor of doublecortin like kinase 1. Nat. Chem. Biol. doi:10.1038/s41589-020-0506-0
- Roskoski, R., Jr. (2020) Properties of FDA-approved small molecule protein kinase inhibitors: A 2020 update. Pharmacol. Res. 152, 104609. doi:10.1016/j.phrs.2019.104609
- Fedorov, O. et al. (2010) The (un)targeted cancer kinome. Nat. Chem. Biol. 6, 166–169. doi:10.1038/nchembio.297
- Edwards, A. M. et al. (2011) Too many roads not taken. Nature 470, 163–165. doi:10.1038/470163a
- Arrowsmith, C. H. et al. The promise and peril of chemical probes. (2015) Nat. Chem. Biol. 11, 536–541. doi:10.1038/nchembio.1867
- Vasta, J. D. et al. (2018) Quantitative, wide-spectrum kinase profiling in live cells for assessing the effect of cellular ATP on target engagement. Cell. Chem. Biol. 25, 206–214 e211. doi:10.1016/j.chembiol.2017.10.010
- Westphalen, C. B. et al. (2017) Functional implication of Dclk1 and Dclk1-expressing cells in cancer. Small GTPases 8, 164–171. doi:10.1080/21541248.2016.1208792
Related Posts
Latest posts by Promega (see all)
- ISO 14001 in Biotech: What It Means for Life Science Researchers - April 2, 2026
- When Cancer Research Depends on Quality RNA: Maxwell® RSC in the Lab - March 30, 2026
- Accelerating Drug Discovery at Grove Biopharma with MyGlo® and ProNect® - December 9, 2025
