Synthetic biology—genetically engineering an organism to do or make something useful—is the central goal of the iGEM competition each year. After teams conquer the challenge of cloning their gene, the next hurdle is demonstrating that the engineered gene is expressing the desired protein (and possibly quantifying the level of expression), which they may do using a reporter gene.
Reporters can also play a more significant role in iGEM projects when teams design their organism with reporter genes to detect and signal the presence of specific molecules, like environmental toxins or biomarkers. Three of the iGEM teams Promega sponsored this year opted to incorporate some version of NanoLuc® Luciferase into their projects.
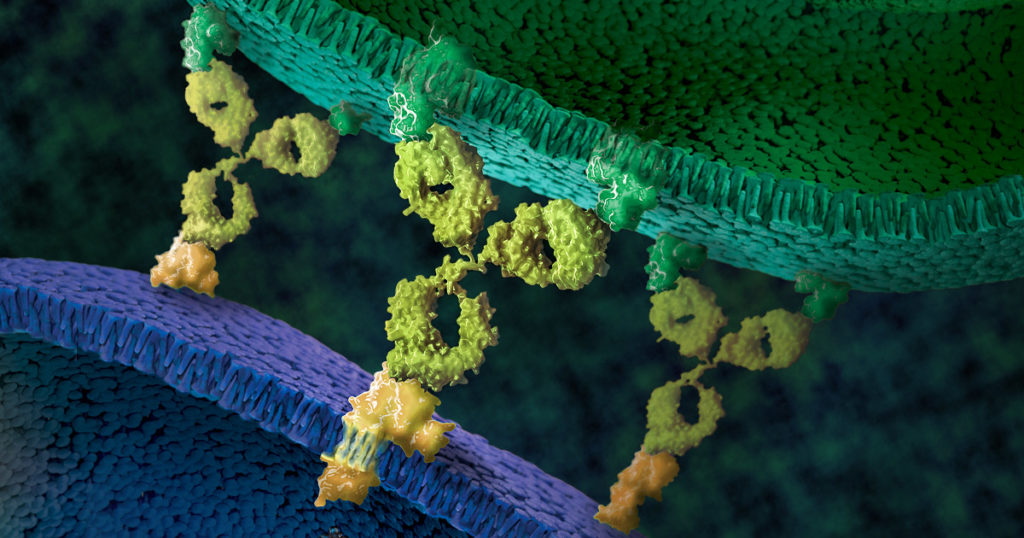
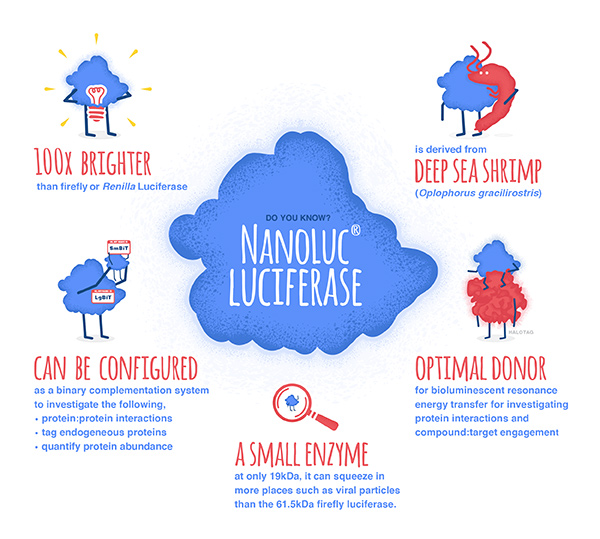
NanoLuc® luciferase is a small monomeric enzyme (19.1kDa, 171 amino acids) based on the luciferase from the deep sea shrimp Oplophorus gracilirostris. This engineered enzyme uses a novel substrate, furimazine, to produce high-intensity, glow-type luminescence in an ATP-independent reaction. Unlike other molecules for tagging and detecting proteins, NanoLuc® luciferase is less likely to interfere with enzyme activity and affect protein production due to its small size.
NanoLuc® Luciferase has also been engineered into a structural complementation reporter system, NanoBiT® Luciferase, that contains a Large subunit (LgBiT) and two small subunit options: low affinity SmBiT and high affinity HiBiT. Together, these NanoLuc® technologies provide a bioluminescent toolbox that was used by the iGEM teams to address a diverse set of biological challenges.
Here is an overview of each team’s project and how they
incorporated NanoLuc® technology.
Continue reading “NanoLuc: Tiny Tag with a Big Impact” Like this:
Like Loading...