Antibiotic-resistant bacteria and their potential to cause epidemics with no viable treatment options have been in the news a lot. These “superbugs,” which have acquired genes giving them resistance to common and so-called “last resort” antibiotics, are a huge concern as effective treatment options dwindle. Less attention has been given to an infection that is not just impervious to antibiotics, but is actually enabled by them.
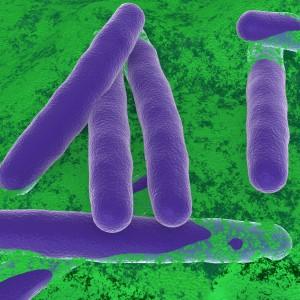
Clostridium difficile Infection (CDI) is one of the most common healthcare-associated infections and a significant global healthcare problem. Clostridium difficile (C. diff), a Gram-positive anaerobic bacterium, is the source of the infection. C. diff spores are very resilient to environmental stressors, such as pH, temperature and even antibiotics, and can be found pretty much everywhere around us, including on most of the food we eat. Ingesting the spores does not usually lead to infection inside the body without also being exposed to antibiotics.
Individuals taking antibiotics are 7-10 times more likely to acquire a CDI. Antibiotics disrupt the normal flora of the intestine, allowing C. diff to compete for resources and flourish. Once exposed to the anaerobic conditions of the human gut, these spores germinate into active cells that embed into the tissue lining the colon. The bacteria are then able to produce the toxins that can cause disease and result in severe damage, or even death.
The ability of C. diff to adhere to the cells lining the walls of the large intestine is a critical factor for developing disease, which has led to research regarding the physiology of this interaction. Of particular interest to researchers is using gene expression analysis to determine what genes are responsible for the characteristics that allow C. diff to adhere to cells lining the gut.
C. diff modulates adherence through the use of proteins that are both present on the cell surface and secreted into the environment. PPEP-1, a metalloprotease, has recently been identified as one of the most highly secreted proteins by C. diff. PPEP-1 is thought to control the shift between adhesion and mobility in these bacteria.
Unfortunately, analyzing gene expression in C. diff is challenging because typical reporters cannot function or are not produced in adequate quantities in the anaerobic environment these bacteria thrive in. Furthermore, no secreted reporters have been developed for C. difficile.
The authors of a 2016 paper published in ACS Synthetic Biology have demonstrated that fusing the signal sequence of PPEP-1 to synthetic constructs produces secreted proteins in C. diff. They then used this approach to create two novel secreted reporters that make it possible to screen gene expression activity in C. diff without requiring complicated sample processing.
One of the reporters created by the researchers is the luciferase reporter sLucopt. It was made by fusing the PPEP-1 signal sequence to a codon-optimized luciferase based on NanoLuc. Luciferase-based bioassays require oxidation in the reaction with the substrate, limiting the use of this reporter with anaerobic organisms like C. diff.
Since the sLucopt reporter is secreted, the culture supernatant can be assayed in an aerobic environment, allowing the original culture to remain anaerobic in order to prevent stress-mediated changes in gene expression. Ultimately, these reporters will make it easier for researchers to perform gene expression analysis in C. difficile by allowing secreted reporters to be measured in aerobic conditions, while eliminating the need for complex sample processing.
Perhaps future studies using reporters such as sLucopt will shine some light on genes that are responsible for the adhesion of C. diff to cells in the colon and lead to an effective treatment option for CDI.
Read the paper:
Oliveira Paiva, A. M. et al. (2016) The Signal Sequence of the Abundant Extracellular Metalloprotease PPEP-1 Can Be Used to Secrete Synthetic Reporter Proteins in Clostridium difficile. ACS Synth Biol. [Epub ahead of print]
Latest posts by Darcia Schweitzer (see all)
- Solving the AAV Titer Challenge: A New Approach to Gene Therapy Workflows - December 10, 2025
- Cytochrome P450 Inhibition: Old Drug, New Tricks - May 5, 2022
- Firefly Luciferase Sheds Light on Development of New Malaria Treatments - April 5, 2021
