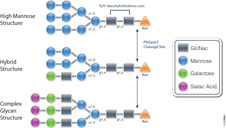
PNGase F (Cat.# V4831) is a recombinant glycosidase cloned from Elizabethkingia meningoseptica and overexpressed in E. coli, with a molecular weight of 36kD.
PNGase F catalyzes the cleavage of N-linked oligosaccharides between the innermost GlcNAc and asparagine residues of high mannose, hybrid, and
complex oligosaccharides from N-linked glycoproteins. PNGase F will not remove oligosaccharides containing alpha-(1,3)-linked core fucose,
commonly found on plant glycoproteins.
Applications
Determining whether a protein is in fact glycosylated is the initial step in glycoprotein analysis. Polyacrylamide gel electrophoresis in the
presence of sodium dodecyl sulfate (SDS-PAGE) has become the method of choice as the final step prior to mass spec analysis. Glycosylated proteins often migrate as diffused bands by SDS-PAGE. A marked decrease in band width and change in migration position after treatment with PNGase F is considered evidence of N-linked glycosylation.
Gel based data are often correlated with information obtained from mass spec analysis. Asn-linked type glycans can be cleaved enzymatically by PNGase F yielding intact oligosaccharides and a slightly modified protein in which Asn residues at the site of de-N-glycosylation are converted to Asp, by converting the previously carbohydrate-linked asparagine into an aspartic acid, a monoisotopic mass shift of 0.9840Da is observed. The deglycosylated peptides are then analyzed by tandem mass spectrometry (MS/MS), and software algorithms are used to correlate the experimental fragmentation spectra with theoretical tandem mass spectra generated from peptides in a protein database.