
Severe acute respiratory syndrome (SARS) is a viral respiratory disease caused by a SARS-associated coronavirus. The most recent version, SARS-CoV-2 was first detected in China in the winter of 2019 and is responsible for the current COVID-19 (coronavirus disease 2019) global pandemic. This virus and its variants have resulted in over 200 million infections and more than 4 million fatalities world-wide. To combat this deadly outbreak the global research community has responded with remarkable swiftness with the development of several vaccines and drug therapies, all produced in record time. In addition to vaccines and drug therapies, diagnostic kits and research reagents continue to roll out to track infections and to help find additional therapies.
This peer-reviewed paper published in Nature Scientific Reports by Alves and colleagues demonstrates how a new assay can be used to discover novel inhibitors that block the binding of SARS-CoV-2 to the human ACE2 receptor as well as study how mutations in the SARS-CoV-2 Spike protein alter its apparent affinity towards human ACE2. The paper also details studies where the assay is used to detect the presence of neutralizing antibodies from both COVID-19 positive samples as well as samples from vaccinated individuals.
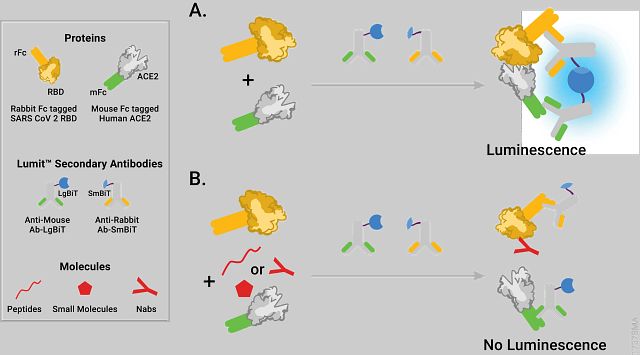
Alves et al. describe development of a bioluminescent assay to measure the interaction of the SARS-CoV-2 Spike Receptor Binding Domain (RBD) protein (rabbit Fc-tagged) with the human Angiotensin Converting Enzyme 2 (ACE2) receptor (mouse Fc-tagged) using the NanoLuc Binary Technology (NanoBiT) technology (see figure). The assay detects the interaction between the rabbit Fc-tagged RBD fragment of SARS-CoV-2 Spike protein and the soluble mouse Fc-tagged ACE2 receptor through binding of LgBiT/SmBiT-conjugated Lumit secondary antibodies.
Learn more about the Lumit™ SARS CoV-2 Spike RBD:hACE2 Immunoassay
The authors discuss how the assay was optimized and present data with known inhibitors of this protein-protein interaction. The inhibitors included specific monoclonal antibodies to SARS-CoV-2 as well as specific peptide inhibitors. Using the Lumit technology they were able to rank order the effectiveness of a range of monoclonal antibodies to inhibit the binding of the Spike RBD protein to hACE2. They also were able to use the assay to characterize a number of Spike RBD mutants and study how the mutations affected the interaction between the mutant proteins and hACE2. The data “showed that the N501Y mutation increased RBD apparent affinity toward ACE2 ten-fold that resulted in escaping inhibition by some anti-RBD antibodies. In contrast, while E484K mutation did not highly change the binding affinity, it still escaped antibody inhibition likely due to changes in the epitope recognized by the antibody.(1)”
They next looked at the affects of neutralizing antibodies from various sources (COVID-19 positive samples as well as samples from vaccinated individuals) and showed that the assay enabled them to assess immune response from the various sources. Thus, the assay can be used for a broad range of applications from drug discovery to clinical research.
Interested in research on viruses? Check out these SARS-CoV-2 and general viral research resources. Or, read our other blogs about SARS-CoV-2 research.
Reference
Alves, J. et al. (2021) A bioluminescent and homogeneous SARS-CoV-2 spike RBD and hACE2 interaction assay for antiviral screening and monitoring patient neutralizing antibody levels. Sci Rep 11, 18428.
Latest posts by Promega (see all)
- Accurate and On-Time: A Look Inside Promega Logistics - June 24, 2025
- Augmenting Human Capabilities with AI Tools - June 17, 2025
- Lead with Empathy: Supporting Caregivers in the Workplace - June 3, 2025
