Have you ever thought about plant viruses? Unless you’re a farmer or avid gardener, probably not. And yet, for many people the battle against agricultural viruses never ends. Plant viruses cause billions of dollars in damage every year and leave millions of people food insecure (1–2), making viruses a major barrier to meeting the United Nations’ global sustainable development goal of Zero Hunger by 2030.
At the University of Western Australia, Senior Research Fellow Dr. Laura Boykin is using genomics and supercomputing to tackle the problem of viral plant diseases. In a recent study, Dr. Boykin and her colleagues used genome sequencing to inform disease management in cassava crops. For this work, they used the MinION, a miniature, portable sequencer made by Oxford Nanopore Technologies, to fully sequence the genomes of viruses infecting cassava plants.
Cassava (Manihot esculenta) is one of the 5 most important calorie sources worldwide (3). Over 800 million people rely on cassava for food and/or income (4). Cassava is susceptible to a group of viruses called begomoviruses, which are transmitted by whiteflies. Resistant cassava varieties are available. However, these resistant plants are usually only protected against a small number of begomoviruses, so proper deployment of these plants means farmers must know both whether their plants are infected and, if so, the strain of virus that’s causing the infection.
Conventional methods for viral identification rely on techniques such as PCR, qPCR, Sanger sequencing and high-throughput sequencing (DNA-Seq and RNA-Seq); however, because samples have to be sent away to labs that perform these analyses, it can take up to 6 months to get sequence data back. MinION uses protein nanopores embedded in a synthetic membrane to sequence DNA. A voltage is applied across the membrane, and as DNA bases are passed through the nanopore proteins, the current changes according to which combinations of bases are in the nanopore. By measuring this current, nanopore sequencers can read native nucleic acid strands without the need for primers—which can introduce bias if the sequence of the primer doesn’t perfectly match the sequence of the DNA.
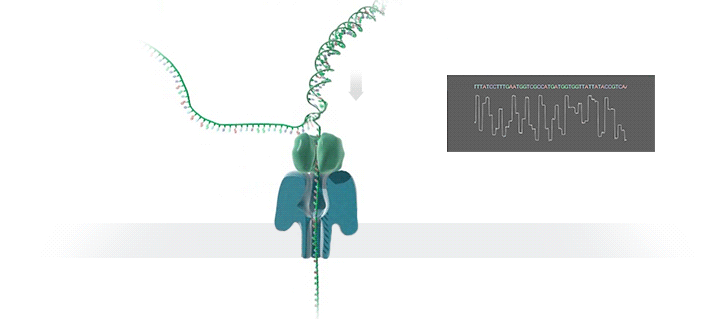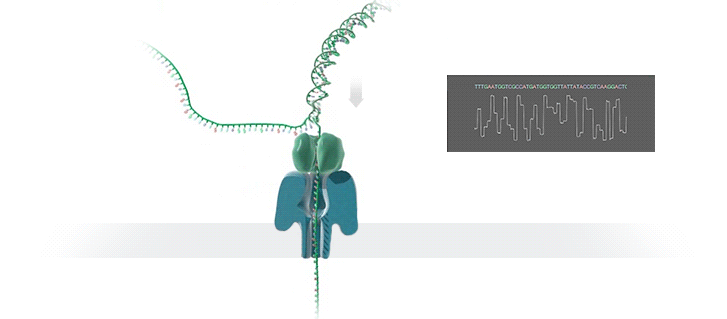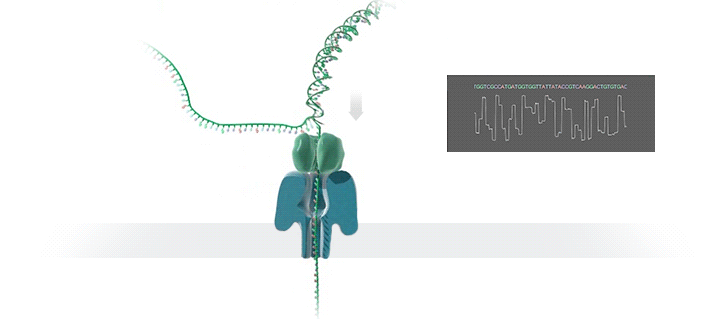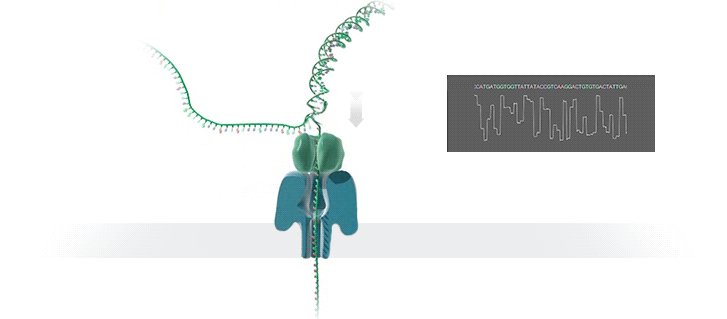
Using the MinION, Dr. Boykin’s team was able to identify what specific strain of virus was infecting a farmer’s plants within 48 hours; in some cases, they have been able to go from sample to answer in just 3 hours (5). The researchers were even able to detect virus in plants that seemed otherwise asymptomatic.
The MinION does come with some challenges: using it requires access to a reliable power source, a computer and the internet, plus the cost of the sequencer would be prohibitive for smallholder farmers. Even so, the advantages—rapid turnaround, minimized sample processing, and the ability to detect mixed infections, latent infections, and even unknown viruses—make this a promising technique for improving plant health regulation and phytosanitation in resource-poor areas.
References
- Subramanya Sastry, K. and Zitter, T.A. (2014) Plant Virus and Viroid Diseases in the Tropics Vol. 2. Springer.
- Legg, J.P. and Thresh, J.M. (2000) Virus Res. 71, 135-49.
- Venturini, M.T. et al. (2016) Agric. 73(6), 520-4.
- Alabi, O.J. et al. (2011) APSnet Features.
- Boykin, L. et al. (2018) F1000 Research 7, 1101.
Latest posts by Julia Nepper (see all)
- Reliable DNA Purification from 3D Cell Cultures - November 18, 2019
- WiSciFest 2019: A Retrospective - October 21, 2019
- Anti-Cancer Drugs Are Pro-Coral - August 26, 2019

