In oncology, tissue biopsies are commonly fixed in formalin and embedded in paraffin (FFPE). These FFPE samples can be used with immunohistochemical or molecular analysis for identifying biomarkers that guide the diagnosis and therapeutic management of patients. This fixation technique allows long-term storage of samples but impacts the integrity of nucleic acids. This makes extracting DNA and RNA from FFPE tissues in sufficient quantity and quality for molecular analysis techniques such as NGS analyses challenging for molecular oncology laboratories.
“At Rennes University Hospital, we receive many lung cancer samples with little material available, or samples of poor quality. The nucleic acid extraction step is therefore critical to get good yield. We have seen that it had a direct impact on the success of downstream analysis,” said Dr. Alexandra Lespagnol. Lespagnol is the Technical Manager of the Molecular Genetics of Cancer core lab at the University Hospital of Rennes in France.

In order to accommodate the increasing number of samples that needed to be analyzed, the Molecular Genetics of Cancer core lab of the University Hospital of Rennes initiated an automation project for extracting DNA from FFPE tissues. The lab also wanted to improve sample tracking and reproducibility of their results.
The lab collaborated with members of the Promega Field Support Scientists team to implement Maxwell® HT DNA FFPE extraction chemistry for processing the samples on a Hamilton STAR liquid handling robot that the lab already owned.
Fast, Consistent and Repeatable Results
First, the Molecular Genetics of Cancer core lab evaluated the performance of the Maxwell® HT DNA FFPE Isolation System using common FFPE biopsy samples and NGS analysis.
“The protocol has been smartly designed and optimized. It is very fast; in less than 2 hours, nucleic acid from 96 samples can be extracted. The protocol is also flexible and can process from 1 to 96 samples with optimal usage of reagents and plasticware,” said Lespagnol.
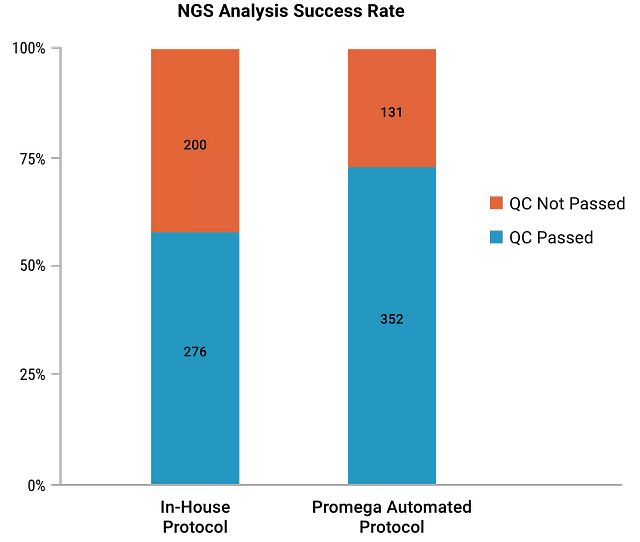
While validating the protocol with NGS analysis, the lab found that the protocol created by the Field Support Scientists with the Maxwell® chemistry led to more than a 15% increase in the NGS success rate compared to an in-house automated protocol. About 500 melanoma, colorectal and lung cancer tumor FFPE tissue samples were tested.
In another test, eight FFPE tissue samples and three blanks were extracted using the kit. No cross-contamination between the samples and the blanks was observed and replicate samples were consistent.
“After we got the first results from samples extracted with the Promega Maxwell® HT FFPE System, we realized that sample quality with our in-house technique was suboptimal. Initially, ovarian tumor samples were tested with the Promega protocol; then we extended its use to all tumor types. Quality improvement is quite phenomenal; we can reduce the number of retesting and reduce time to results,” said Lespagnol.
The Molecular Genetics of Cancer core lab plans to expand automated extraction protocols to other sample types, including liquid biopsy (circulating cell-free DNA from plasma) and RNA from FFPE. Testing for these applications has already begun.
Automation Experts Ready to Help
This project was carried out with the collaboration and assistance of the Promega Field Support Scientists (FSS) team, especially Nans Bodet. The FSS team are experts in automating Promega chemistries for nucleic acid extraction, quantification and amplification, as well as cell-based assays on many types of liquid-handling platforms. The team supports every aspect needed to implement automation, from developing specific scripts to on-site implementation, staff training and ongoing assistance with new projects.
“[The] Promega team perfectly understood our needs and translated it into solutions,” said Lespagnol.
Learn more about high-throughput automated nucleic acid extraction solutions and our team of Field Support scientists here.
Latest posts by Jordan Nutting (see all)
- The Central Dogma of Promega: The Story and Science Behind Our Kit Packaging Design - May 7, 2024
- Silencing the Immunogenicity of AAV Vectors - April 4, 2024
- Discovering Cyclic Peptides with a “One-Pot” Synthesis and Screening Method - February 29, 2024
