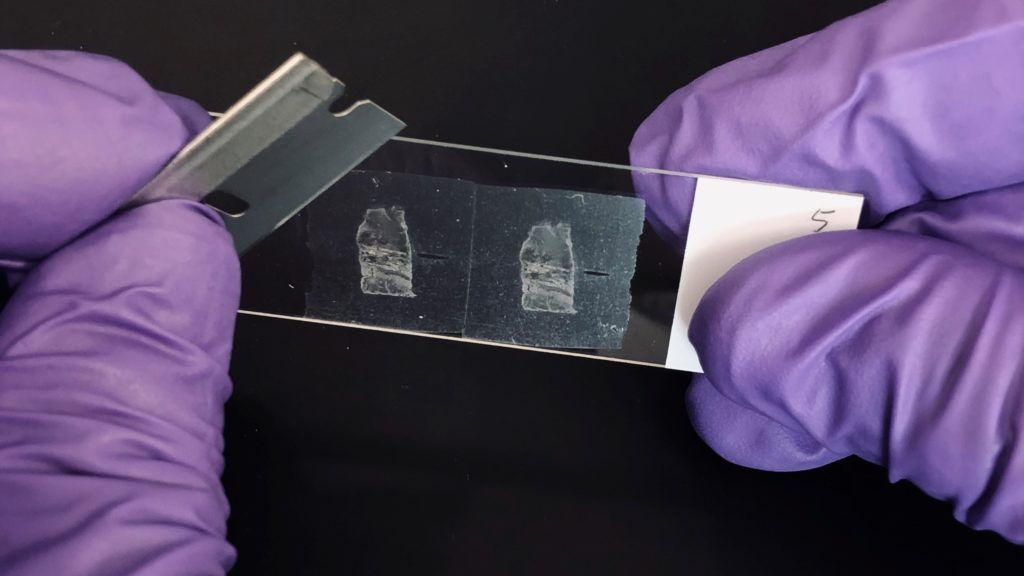
Reliable molecular research starts with reliable sample preparation. Two recently published cancer biology studies illustrate this well, and both studies relied on the Maxwell® RSC platform to extract RNA from formalin-fixed, paraffin-embedded (FFPE) tissue, the archival format that makes up the bulk of clinical pathology material.

Mapping Molecular Targets in a Rare Thyroid Cancer
A 2025 study published in Endocrine Pathology focused on poorly differentiated thyroid carcinoma (PDTC), a rare and aggressive thyroid cancer subtype with limited treatment options once surgery is no longer curative (1). The research question was straightforward but clinically urgent: how many PDTC cases harbor mutations that could be targeted with existing or emerging therapies?
Continue reading “When Cancer Research Depends on Quality RNA: Maxwell® RSC in the Lab”