The polymerase chain reaction (PCR) has revolutionized modern biology as a quick and easy way to generate amazing amounts of genomic data. However, when PCR doesn’t work, it can be frustrating. At these times, PCR and reverse transcription PCR (RT-PCR) inhibitors seem to be everywhere: They lie dormant in your starting material and can co-purify with the template of interest, and they can be introduced during sample handling or reaction setup. The effects of these inhibitors can range from partial inhibition and underestimation of the target nucleic acid amount to complete amplification failure. What is a scientist to do?
The first step is education; you must learn what you’re up against. Here, I will focus on signs that inhibitors are causing problems with your quantitative PCR.
Before you can overcome the adverse effects of PCR inhibitors, you must realize that there is a problem. Sounds easy, right? Either the PCR worked or it didn’t. Unfortunately, it’s not that simple. PCR and RT-PCR inhibition often is not an all-or-nothing phenomenon and can have a number of different causes, including inhibition of the DNA polymerase or reverse transcriptase, chelation of necessary co-factors and interference with primer binding as well as causes that mimic true inhibition such as adsorption of the template to the tube, template or primer degradation and interference with the fluorescent signal.
For qPCR, often you can learn something about the type of inhibition from shifts in the quantification cycle (Cq, previously known as CT) value, the point at which the amplification curve crosses a detection threshold where signal due to product accumulation can be distinguished reliably from background. The Cq value is inversely proportional to the amount of template: the more template that is present, the earlier the target is detected.
You can also learn something from the slope of a graph created by plotting log nucleic acid concentration versus Cq value for a series of nucleic acid dilutions. For a qPCR with 100% efficiency, the slope of such a graph will be –3.32. A slope of –3.3 to –3.6 indicates good amplification efficiency (100% to 90%, respectively), and a slope below –3.6 indicates poor efficiency and possible inhibition.
Here are just a few examples of how qPCR inhibition can be detected by examining the data and amplification curves.
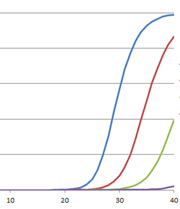
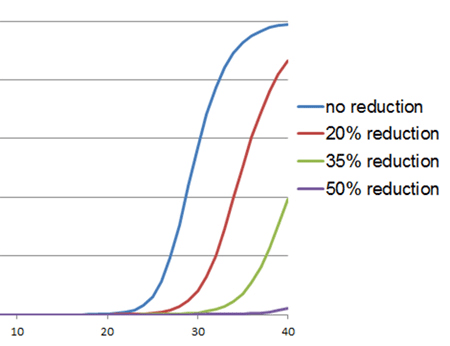
Inhibition of Taq DNA polymerase is characterized by an increase in Cq value and a change in amplification curve morphology, as shown in the example to the right. However, increased Cq values also may indicate lower amounts of target. This possibility can be distinguished from an increased Cq value due to inhibition of Taq DNA polymerase through the use of an internal PCR control (IPC). The IPC is a set of primers and template that is included in the reaction mix along with specific primers for the primary assay targets. Amplification of the IPC should occur with equal efficiency (similar Cq values) in all samples; if the IPC Cq value of the test sample is substantially delayed relative to that of a standard or positive control, then this indicates an inhibitor in the test sample. Another way to make this distinction is to serially dilute the template and look for smaller-than-expected shifts in Cq values: At 100% efficiency, the amount of product doubles after each cycle. Thus, for tenfold dilutions, the Cq values should be approximately 3.3 cycles apart (23.3 = 10); a Cq shift of less than 3.3 cycles would strongly suggest inhibition.
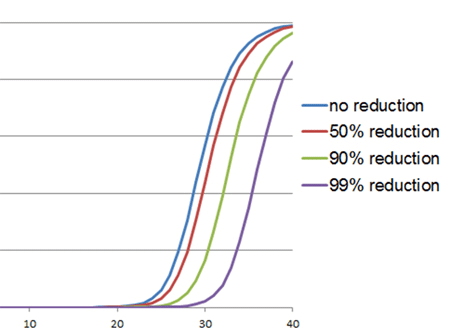
In RT-qPCR, inhibition of the reverse transcriptase is characterized by a shift in the Cq values. The amplification curve will look perfectly normal (as shown to the right). This type of inhibition will result in underestimation of the target RNA amount in the starting material and cause the Cq value to be shifted to the right. Again, the IPC and serial dilutions of the template can be used to determine whether this shift is due to inhibition or simply a lower amount of target.
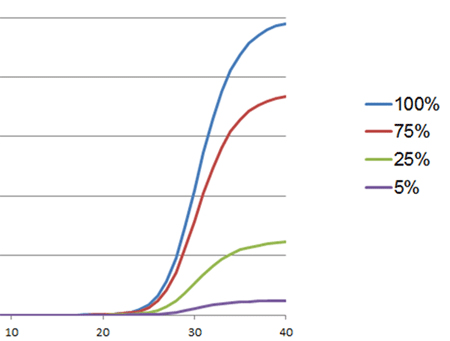
Another reason that qPCR appears to fail is interference with the fluoresence signal. This interference can occur when an inhibitor competes with the amplicon for binding to the fluorescence dye used for detection (e.g., BRYT Green® or SYBR® Green) or when the inhibitor interacts with the fluorescently labeled probe. Typically, background fluorescence increases, and specific fluorescence decreases. As a result, the amplification curves appears flattened compared to amplification curves of uninhibited reactions. In severe cases of inhibition, the amplification curve never crosses the threshold, and a Cq value is not called.

excellent attempt!
I have one Inhibitor that aways makes the first point of the dilution curve amplify after the others more diluted. That makes sense? Any idea what it is. They are RNA from adipose tissue, with good ratios and profile in the gel
Hi Marilla, I will refer this question our technical services scientists and ask them to reply. Thanks, Michele
If you are asking if it makes sense that the undiltued sample with inhbitors amplifies later than the diluted one makes sense, then yes. That does make sense. Think generically of a dose-response curve for an inhibitor of any enzyme, where relatively small changes in concentration of the inhibitor can have drastically different effects (or not) on the enzyme. Often nearly the full range of effect of the inhibitor may be observed within ~1 order of magnitude. As for the source of your inhibitor: one possibility is a component of the RNA extraction workflow, e.g., a guanidine compound, if it was carried over. The precise inhibitor may not be easily identifiable, but the source, and strategy for mitigation, may be approachable. Please let us know if you have additional questions, and always feel free to contact technical services for troubleshooting help. Email: techserv@promega.com or chat with them during US hours https://www.promega.com/support/