This blog was written by guest contributor Tian Yang, Associate Product Manager, Promega, in collaboration with Kristin Huwiler, Manager, Small Molecule Drug Discovery, Promega.
During the development of chemical probes or small-molecule drugs, compound affinity (Kd) or potency (IC50) is used to characterize compound-target interactions, to guide structure-activity relationship analysis and lead optimization and to assess compound selectivity.
However, neither parameter provides information on how quickly a compound engages with and dissociates from the target. The dissociation constant Kd reflects the relative concentrations of unbound and bound state of the compound at thermodynamic equilibrium, and while IC50 is an empirical metric that measures the concentration at which a biochemical or cellular process is reduced to half of the maximum value, IC50 values are typically determined when the process is assumed to be at equilibrium or steady-state. For a closed system, like cells in a culture dish, these thermodynamic parameters are quite informative. In an open system like the human body, where compound-target interactions often do not reach equilibrium, the kinetic parameters, in addition to the thermodynamic parameters, are needed to better understand and characterize compound target engagement over time (1,2).
For a basic binary compound-target interaction, the binding kinetics can be defined by the association rate constant (kon) and the dissociation rate constant (koff). The latter is often expressed as residence time, which is 1/koff and describes the length of time a compound remains bound to its biological target(s). For compounds that are removed rapidly in an open system, optimizing compound residence time in particular can help deliver sustained activity for target modulation, which improves compound efficacy and allows for greater target selectivity mediated by differences in binding kinetics. Specifically, under non-equilibrium conditions, targets with long residence time and low turnover rate will be selectively engaged over targets with short residence time and/or high turnover rate. This kinetic selectivity can occur even when targets exhibit similar compound affinity under thermodynamic equilibrium (i.e., targets with similar Kd or IC50), providing an additional avenue to achieve on-target specificity and minimize off-target interactions.
Methods for Measuring Residence Time
Compared to methods for characterizing compound binding affinity, techniques for assessing kinetic parameters of compound binding can be more laborious (1). Nevertheless, several methodologies have been developed to measure compound residence time, including methods that allow for higher throughput.
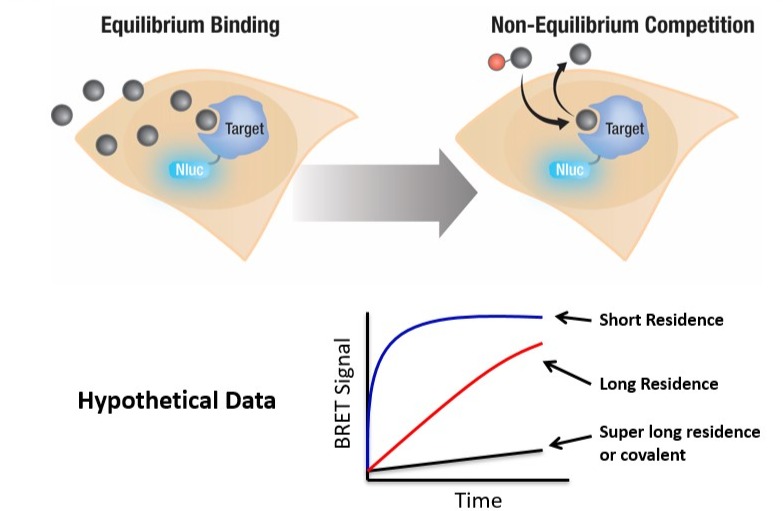
Radio- or fluorescently-labeled compounds, which are frequently used to determine compound binding affinity, can also be applied to assess compound residence time (3). The label can be introduced to the compounds of interest to directly monitor compound association or dissociation. Alternatively, a probe-displacement approach can be used, where unlabeled compounds compete with a labeled probe molecule for target binding, and the kinetic rates can be determined using the Motulsky & Mahan method or the compound washout method (3). In addition, the fluorescent probes used in Fluorescence Polarization, Time Resolved-Fluorescence Resonance Energy Transfer, or Bioluminescence Resonance Energy Transfer (BRET) techniques only generate desired signals when the probe is bound to the target, removing the need to separate unbound probes and thereby making these techniques more suitable for higher-throughput applications.
As an alternative to the labeled ligand approaches, Surface Plasmon Resonance (SPR) is a probe-free technique that has been commonly used for studying compound binding kinetics (3). In SPR, the target is immobilized on a thin gold film within a flow cell. As ligand-containing solutions flow across the surface, binding events change the local refractive index near the sensor in real-time, which can be detected as a signal for compound binding. By monitoring the SPR signal in the association phase and the dissociation phase, precise determination of kon and koff can be made, making SPR well-suited for characterizing kinetic parameters of compound-target interactions (3,4). In addition to SPR, bio-layer interferometry and isothermal titration calorimetry are also probe-free methods that can be used for kinetic analysis of compound binding.
Although these methods are often used with purified targets in biochemical assays, some have been adapted for analyzing compound residence time in live cells. For example, while SPR is typically performed with immobilized proteins, strategies have been developed to immobilize cells or ligands of interest to measure the interaction between cell-surface protein and its ligand in real-time (5). NanoBRET, a BRET-based technique for measuring molecular proximity in live cells, has been applied to measure compound-target interactions in live cells and is capable of kinetic analysis to assess compound residence time for both intracellular and extracellular targets (6,7).
Efficacy, Toxicity and Residence Time
Compound affinity and potency measurements, which are determined under equilibrium or at steady-state, are often used during drug development to guide lead optimization and improve compound efficacy to achieve the desired biological outcomes. However, in a non-equilibrium system, compound residence time, instead of affinity, can be the determining factor for efficacy (8). In their 2021 publication, Lyczek and colleagues investigated the mechanism of imatinib-resistance mediated by Bcr-Abl mutations that are distal from the drug-binding site (9). Among the mutants they assessed, about one third of the mutants displayed a reduced residence time for imatinib, with some dissociating twice as fast.
Interestingly, a couple of the imatinib-resistant mutants had significantly reduced residence time despite exhibiting similar or even improved affinity for imatinib, suggesting that drug resistance could be mediated by decreasing drug residence time in addition to reducing drug binding affinity.
Just as residence time can affect compound efficacy, compounds can display kinetic selectivity by having different residence times for related targets. Typically, compound selectivity is determined by comparing the affinity, potency or target enrichment for a compound among a panel of potential targets, which are based on equilibrium binding experiments. However, this thermodynamic selectivity does not always correlate with compound selectivity based on binding kinetics, and kinetic selectivity can exist even when a compound does not show thermodynamic selectivity (2).
As an example, when ponatinib, a Bcr-Abl inhibitor, was profiled against a group of 10 kinases using SPR, ponatinib inhibited 6 kinases, including Abl, with similar potency based on enzyme activity assays, which would suggest that ponatinib is non-selective among these kinases. In addition, 3 kinases were inhibited with greater potency by ponatinib than Abl (10). However, when comparing compound residence time, ponatinib preferentially engages Abl with a residence time of ~180 hours, which is more than 25 times longer than the kinase with the second longest residence time. In an in vivo system where the compound can be readily eliminated, the long residence time of ponatinib at Abl can ensure persistent inhibition of Abl, while the activity of other kinases could recover upon dissociation of ponatinib following compound elimination from the system. This difference in residence time can enable a more selective inhibition of Abl that would not appear possible based on the compound potency measurements.
Beyond Binary Complexes – Residence Time and Targeted Protein Degradation
Selectively targeting proteins for removal from the cell, instead of directly modulating protein activity, has emerged as a new modality for therapeutic development. The protein is targeted for degradation using the cell’s natural ubiquitin proteasome system, and this process is initiated by compounds, such as molecular glues and proteolysis targeting chimeras (PROTACs), that bring the target protein and an appropriate E3 ligase into a functional ternary complex. Consequently, how well a compound can induce ternary complex formation can significantly impact the degradation outcome.
In a recent publication by Fan et al., the authors took a more systematic investigation of how degrader residence time at target protein can affect target protein degradation (11). Using a series of pan-kinase binder analogues with varying rates of dissociation but similar binding profiles, Fan and colleagues generated corresponding PROTACs to probe how differences in residence time can impact target protein degradation across numerous kinases. To validate that the PROTACs engaged the target kinase similarly to the parent compounds, a NanoBRET® Target Engagement assay was performed to assess cellular residence time, and the results matched the expectations. With these tool compounds in hand, the authors assessed target occupancy, ternary complex formation, target ubiquitination and target degradation induced by each compound. They found that different kinases displayed varying preferences of target residence time to achieve optimal degradation, agreeing with the predictions of mathematic models that optimal degradation is dependent on the balance of ternary complex cooperativity and target residence time. Although the data did not show correlation between optimal residence time and experimental or predicted scores of degradability, the authors found that changes to target residence time can significantly impact degradation efficacy and selectivity of the degrader compounds, and a systematic evaluation of off-rate in early stage of degrader designs could facilitate degrader development and optimization.
Summary
Residence time, defined as the duration a drug remains bound to its biological target, is increasingly recognized as a critical parameter in drug discovery and can profoundly impact a compound’s efficacy and specificity. Although residence time has been more challenging to characterize historically, recent methodology advances have simplified the accurate measurement of residence time, both in biochemical assays and in live cells, making it possible to routinely incorporate kinetic analysis into the evaluation of target engagement. By combining residence time analysis with the traditional measures of affinity and potency for target engagement, researchers can attain a more comprehensive understanding of compound-target interactions. This broader view can uncover hidden selectivity, explain unexpected resistance mechanisms, and guide the design of drugs with better therapeutic potential.
References
- Srinivasan, B. (2023) A guide to enzyme kinetics in early drug discovery. FEBS J. 290(9), 2292-2305.
- Tonge, P.J. (2018) Drug–Target Kinetics in Drug Discovery. ACS Chem. Neurosci. 9(1), 29-39.
- Bernetti, M., et al. (2019) Kinetics of Drug Binding and Residence Time. Annu. Rev. Phys. Chem. 70, 143-171.
- Georgi, V., et al. (2017) Binding kinetics in drug discovery – A current perspective – PubMed Front. Biosci. (Landmark Ed). 22(1), 21-47.
- Mamer, S.B., et al. (2019) The Convergence of Cell-Based Surface Plasmon Resonance and Biomaterials: The Future of Quantifying Bio-molecular Interactions—A Review. Ann. Biomed. Eng. 48(7), 2078-2089.
- Robers, M.B., et al. (2015) Target engagement and drug residence time can be observed in living cells with BRET. Nat. Commun. 6, 10091.
- Boursier, M.E., et al. (2020) The luminescent HiBiT peptide enables selective quantitation of G protein-coupled receptor ligand engagement and internalization in living cells. J. Biol. Chem. 295(15), 5124-5135.
- Copeland, R.A. (2016) The drug–target residence time model: a 10-year retrospective. Nat. Rev. Drug Discov. 15, 87-95.
- Lyczek, A., et al. (2021) Mutation in Abl kinase with altered drug-binding kinetics indicates a novel mechanism of imatinib resistance. Proc. Natl. Acad. Sci. U.S.A. 118(46), e2111451118.
- Willemsen-Seegers, N., et al. (2017) Compound Selectivity and Target Residence Time of Kinase Inhibitors Studied with Surface Plasmon Resonance. J. Mol. Biol. 429, 574-586.
- Fan, A.T., et al. (2025) A Kinetic Scout Approach Accelerates Targeted Protein Degrader Development. Angew. Chem. Int. Ed. 64, e202417272.
Latest posts by Promega (see all)
- Accelerating Drug Discovery at Grove Biopharma with MyGlo® and ProNect® - December 9, 2025
- From Mt. Fuji to the Lab Bench: A UW-Madison Student’s Summer in Japan - December 3, 2025
- Promega Fc Effector Assays: Measure Every Mechanism - November 26, 2025
