mRNA-based therapeutics are being explored across a range of applications, including vaccines, protein replacement and immunotherapies (2).
Before any formulation decisions enter the picture, teams need confidence in the RNA itself: that it is the right sequence, right properties and the right purity to behave predictably downstream. That is where it helps to separate drug substance from drug product. The drug substance is the active ingredient intended to deliver a pharmacological effect, while drug product is the finished dosage form that contains that ingredient (6).
This post focuses on what happens upstream, making the mRNA drug substance before formulation. In practical terms, that upstream work spans choosing an mRNA construct, producing it by IVT, and then purifying and analyzing the product so it has the desired quality attributes (5).
What is an mRNA construct
An mRNA construct is the engineered RNA sequence a team advances because it’s expected to express a therapeutic protein (or deliver another therapeutic effect; 2). At this stage it’s still a design, but the goal is to turn that design into drug substance: a reproducible, well-characterized mRNA material you can carry forward. In early development, construct can sound abstract, but the practical question is: can you repeatedly produce the same mRNA you designed, at a high-quality level.
The high-level science behind IVT
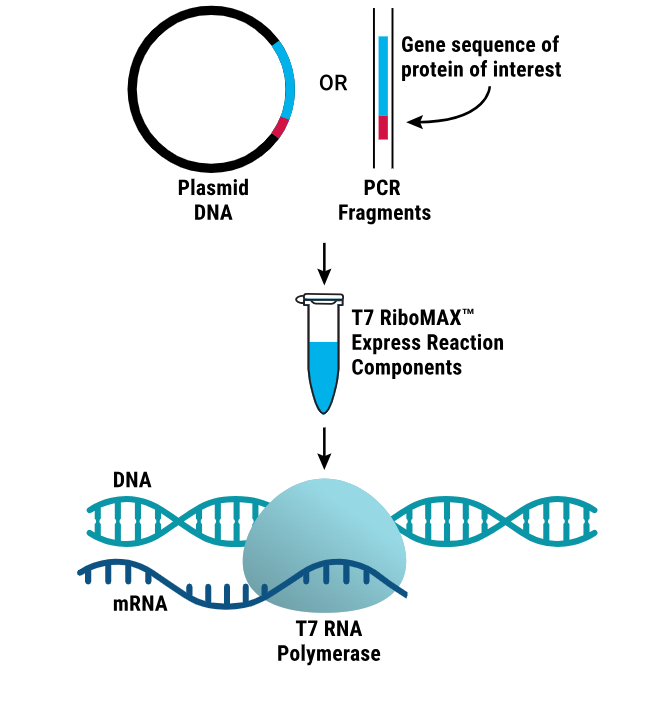
A common way to generate mRNA in development workflows is using in vitro transcription (IVT) which is a template-directed, cell-free process for creating mRNA. At a high level, IVT can be thought of as combining a DNA template with ribonucleotides and an RNA polymerase to synthesize RNA (2,4). IVT is commonly performed with bacteriophage RNA polymerases such as T7, T3 or SP6, depending on the promoter built into the template (3,4).
IVT depends on thoughtful template engineering: the DNA template includes a promoter sequence (for example, T7) upstream of the sequence of interest, 5’ and 3’ untranslated regions and a poly(A) tail (4).
A simple map for making mRNA
- Build the template you intend to transcribe
- Template format shapes what the polymerase can produce and how cleanly you can interpret the result. Common template types include linearized plasmids, PCR products that incorporate a promoter sequence, or annealed oligonucleotide templates (often for shorter RNAs; 4).
- Design the IVT reaction (conceptually)
- Most IVT workflows rely on the same core categories of inputs: a polymerase, rNTPs and supporting reagents that protect the RNA and support reaction performance. Examples of common categories include T7 RNA polymerase, RNase inhibition and inorganic pyrophosphatase (4,8,10). For teams comparing options, Promega also includes a side-by-side comparison of high-yield IVT kits, which can be a helpful starting point when selecting a system for your RNA production needs (5).
- Decide how you want the RNA to behave downstream
- Depending on the target application, teams may choose capped or uncapped RNA and standard or modified nucleotides (5). At a high level, these choices can influence readiness for translation, consistency of output and analytical characterization aspects. Cap analogues can also be used during transcription initiation to generate more native-like 5′ ends that support translation (2,4).
These are the main upstream decision points, but typical workflows also include template removal, purification and basic characterization before formulation (5).
Explore Flexible, Scalable RNA Production Solutions

Our RiboMAX™ mRNA production solutions offer high-yield in vitro transcription systems with cGMP and animal origin-free IVT reagents options. When paired with Promega scalable, automation-friendly purification technologies, our IVT systems enable, high-quality mRNA production.
Sources of variability in IVT mRNA manufacturing
One of the most common frustrations in mRNA synthesis is getting material that looks fine at first glance but carries impurities or heterogeneity that show up later as variability, weak translation, or confusing readouts.
There are many variables that affect yield and mRNA quality, but a few recurring issues tend to show up across workflows, including manufacturing variables, purification tradeoffs and scale sensitivity:
- Sequence and construct design
- Variability often starts at the sequence and transcript-design level. Factors like codon usage, sequence composition (including GC content) and structural elements can influence transcription and how consistently full-length product is generated (4). Transcript-end design choices also matter, including the 5’ end, 3’ tailing approach and UTR selection, since these decisions can affect downstream translation performance and what needs to be verified during characterization (2,4).
- Reaction design and inputs
- Reaction setup matters too: polymerase selection, standard vs modified nucleotides and supporting reagents used to protect RNA and support reaction performance can shift the yield and consistency from run to run (4). Because the IVT reaction environment influences both product integrity and byproduct formation, teams often treat reaction optimization as part of the same manufacturing story as purification rather than a standalone step (4).
- Cleanup and purification
- After transcription, the next task is straightforward in concept: remove what you don’t want carried forward. This, however, can be much more difficult in practice when you are balancing recovery with consistency. Cleanup separates the mRNA from leftover reaction components like enzymes, nucleotides, template DNA and dsRNA so they don’t interfere with downstream characterization or functional readouts (4,5). Our IVT cleanup resource describes purification of mRNA from IVT reactions as a defined step in the workflow. In many methods, mRNA is captured on a column or beads, washed and then eluted into a small volume (7).
- Scale changes the rules
- Many workflows that look stable in a single tube become more sensitive as volume and throughput increase. Even at the early manufacturing stage done in smaller scales, scalability is often treated as a first-class constraint, not a future problem. Planning for scale early often means standardizing inputs and documenting conditions so results stay comparable as volume and throughput change (5,10).
Where animal origin–free fits
Animal origin–free (AOF) and cGMP-manufactured inputs matter for a practical reason: they reduce avoidable variables. Animal-derived raw materials can introduce added variability and traceability/risk-assessment burden (including TSE/BSE considerations), so AOF inputs help reduce both scientific and regulatory unknowns early (8,10).
As programs mature, teams often place more emphasis on input consistency, traceability, and documentation. Using well-characterized IVT reagents and recording lot-to-lot performance can help reduce avoidable variability as workflows scale and as requirements become more formal (8,10).
Interested in more about animal-origin-free IVT? See the RiboMAX™ AOF RNA Production System page (3,5).
Conclusion
In many programs, downstream challenges can trace back to upstream foundations. When manufacturing the mRNA is defined clearly with rational template design, quality IVT reagents, purification and early checks, it becomes easier to troubleshoot, scale and maintain comparability as the work progresses (4,5). Even early on, screening for length/heterogeneity and key impurities like dsRNA can help avoid confusing downstream readouts (4,5).
If you’re exploring upstream workflow options, our mRNA Therapy Solutions materials highlight support for RNA synthesis, purification and characterization across development stages (5,10).
References
- Beckert, B., & Masquida, B. (2010). Synthesis of RNA by in vitro transcription. RNA, 703, 29–41. https://doi.org/10.1007/978-1-59745-248-9_3
- Garber, K. (2022). mRNA pioneers refocus on therapeutics. Nature Reviews Drug Discovery, 21, 699–701. https://www-nature-com.ezprom01.infotrieve.com/articles/d41573-022-00156-5
- In Vitro Transcription | RNA Synthesis Kits | RNA Production. (2026). Promega.com. https://www.promega.com/products/rna-analysis/in-vitro-transcription/
- Lenk, R., Kleindienst, W., Szabó, G. T., Baiersdörfer, M., Boros, G., Keller, J. M., Mahiny, A. J., & Vlatkovic, I. (2024). Understanding the impact of in vitro transcription byproducts and contaminants. Frontiers in Molecular Biosciences, 11. https://doi.org/10.3389/fmolb.2024.1426129
- mRNA Therapy Solutions. (2026). Promega.com. https://www.promega.com/applications/gene-therapy-tools/rna-therapy/mrna-therapy-solutions/
- Patel, P. R. (2023). Chemistry, Manufacturing and Controls: Regulatory Considerations Through Clinical Development. https://www.fda.gov/media/175396/download
- Promega Corporation. (2024, June). IVT RNA ProNex [Application note] (Publication No. PA1001; Version 06/24). https://promega.widen.net/s/hchgrkqvrm/ivt-rna-pronex-pa1001
- Promega Corporation. (2024, October). Custom raw materials for mRNA synthesis [Flyer] (Publication No. FL682I). https://promega.widen.net/s/jvwcfpprrd/custom-raw-materials-for-mrna-synthesis-a4-fl682i-print
- Promega Corporation. (2025, October). In vitro transcription [Flyer] (Publication No. GFN368). https://promega.widen.net/s/rpss2spqkz/ivtranscription_ltr_gfn368
- Raw Materials for mRNA Therapeutic Manufacture. (2026). Promega.com. https://www.promega.com/custom-solutions/custom-manufacturing/mrna-manufacturing/
Discover Characterization and QC Assays for mRNA

Our comprehensive portfolio of mRNA analytical characterization and QC solutions enable sensitive, quantitative assessment of mRNA quality. We offer assays for dsRNA impurity detection, accurate RNA quantitation, and cell-based potency determination to ensure safety, consistency and performance of mRNA therapeutics.
Latest posts by Shannon Sindermann (see all)
- Some St. Patrick’s Day Science: Green Rivers, Four-Leaf Clovers and Optics of a Good Pint - March 17, 2026
- Sequence to Substance: Making the mRNA Therapeutic - February 17, 2026
- The Science of Slipping… Blame the Molecules! - January 29, 2026
