Our second installment of the Promega qPCR Grant Recipient blog series highlights Dr. Laura Leighton, a trained molecular biologist and postdoctoral researcher at the Australian Institute for Bioengineering and Nanotechnology. Leighton’s scientific journey features a passion for molecular biology and problem-solving. Her path has been illuminated by mentorship, relationships with fellow scientists and a commitment to creativity in overcoming challenges. Here, we explore her scientific journey, reflect on research lessons and foreshadow her plans for the Promega qPCR grant funds.
Dr. Laura Leighton grew up in a rural area in Far North Queensland, Australia, where she spent her early life exploring critters on the family farm. Her upbringing was infused with a deep connection to the environment, from raising tadpoles in wading pools to observing wildlife and witnessing food grow firsthand. Observing the biology around her ultimately piqued her interest in science from a young age. She then began her academic journey in 2011 at the University of Queensland, Australia. She studied biology while participating in a program for future researchers, which led her to undergraduate research work in several research labs. She dabbled in many research avenues in order to narrow in on her scientific interests all while adding different research tools to her repertoire.
After serving as a research assistant in Dr. Timothy Bredy’s lab, she decided to continue work in this lab and pursue a PhD in molecular biology. During her PhD, Leighton worked on several projects from cephalopod mRNA interference to neurological wiring in mice. The common thread in these projects is Leighton’s passion for the puzzles of molecular biology:
“I also love molecular engineering and the modularity of molecular parts. There’s something really special about stringing together sequence in a DNA editor, then seeing it come to life in a cell,” she says.
Continue reading “Promega qPCR Grant Series #2: Molecular Biologist, Laura Leighton”

 Let’s face it, most lab techs and purchasing agents aren’t all that happy when you send them an Instagram picture of your latest lunchroom-napkin cloning strategy as your order form for your next big cloning experiment. So we have created the
Let’s face it, most lab techs and purchasing agents aren’t all that happy when you send them an Instagram picture of your latest lunchroom-napkin cloning strategy as your order form for your next big cloning experiment. So we have created the 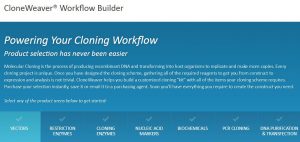 Already have a favorite vector and a freezer full of restriction enzymes? No problem, skip those steps and move on to getting the perfectly sized nucleic acid markers or the particular polymerase your experiment requires.
Already have a favorite vector and a freezer full of restriction enzymes? No problem, skip those steps and move on to getting the perfectly sized nucleic acid markers or the particular polymerase your experiment requires.