It is with sadness that we recognize the passing of Dr. Frederick Sanger. Sanger is known to molecular biologists and biochemists worldwide for his DNA sequencing technique, which won for him the 1980 Nobel prize in Chemistry.
Also noteworthy, Sanger’s laboratory accomplished the first complete genome sequence, that of a viral DNA genome more than 5,000 base pairs in length.
The 1980 prize was Sanger’s second Nobel award, his first awarded in 1958 for determining the chemical structure of proteins. In this work, Sanger elucidated not only the amino acids that comprised insulin but also the order in which the amino acids occurred.
About Sanger Sequencing
Sanger DNA sequencing is also known as the chain-termination method of sequencing. The Sanger technique uses dideoxynucleotides or ddNTPs in addition to typical deoxynucleotides (dNTPs) in the reaction. ddNTPs result in termination of the DNA strand because ddNTPs lack the 3’-OH group required for phosphodiester bond formation between nucleotides. Without this bond, the chain of nucleotides being formed is terminated.
Sanger sequencing requires a single-stranded DNA, a DNA primer (either radiolabeled or with a fluorescent tag), DNA polymerase, dNTPs and ddNTPs. Four reactions are set up, one for each nucleotide, G, A, T and C. In each reaction all four dNTPs are included, but only one ddNTP (ddATP, ddCTP, ddGTP or ddTTP) is added. The sequencing reactions are performed and the products denatured and separated by size using polyacrylamide gel electrophoresis.
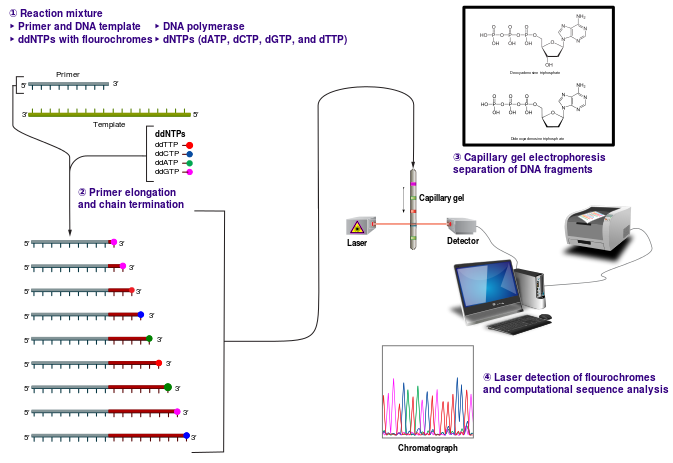
This reaction mix results in various lengths of fragments representing, for instance, the location of each A nucleotide in the sequence, because while there is more dATP than ddATP in the reaction, there is enough ddATP that each ATP ultimately instead is replaced with a ddATP, resulting in chain termination. Separation by gel electrophoresis reveals the size of these ddATP-containing fragments, and thus the locations of all A nucleotide in the sequence. Similar information is provided for GTP, CTP and TTP.
The Maxam and Gilbert DNA sequencing method had the advantage at the time of being used with double-stranded DNA. However, this method required DNA strand separation or fractionation of the restriction enzyme fragments, resulting in a somewhat more time-consuming technique, compared to the 1977 method published by Sanger et al.
Dr. Sanger was born in Gloucestershire, U.K. in 1918, the son of a physician. Though he initially planned to follow his father into medicine, biochemistry became his life-long passion and area of research endeavor. Sanger retired at age 65, to spend more time at hobbies of gardening and boating.
References
Sanger, F. , Nicklen, S. and Coulson, A.R. (1977) DNA sequencing with chain-terminating inhibitors. Proc. Natl. Acad. Sci. USA 74, 5463-7.
Maxam, A.M. and Gilbert, W. (1977) A New Method for Sequencing DNA. Proc. Natl. Acad. Sci. USA
There is something special about seeing the original Sanger publication from 1977, available here as a scan.
Kari Kenefick
Latest posts by Kari Kenefick (see all)
- Fluorescent Ligands in Biological Research: Where We’ve Been, Where We’re Headed - June 27, 2024
- Cell-Based Target Engagement and Functional Assays for NLRP3 Inhibitor Profiling Help Identify Successes and Failures - January 25, 2024
- Illuminating the Brain with a New Bioluminescence Imaging Substrate - April 13, 2023

a nice one,….this keeps one remembering the classic days….!
Inspiring to read of the accomplishments of those on whose shoulders we stand.
Thanks for your comment, Bob. What you say is so true!
prof premraj pushpakaran — 2018 marks the 100th birth year of Frederick Sanger!!!
Thanks for letting us know about this important anniversary this year.