Today’s blog is guest-written by Wihan Adi, a Master’s student majoring in physics at Justus-Liebig-University in Giessen and team member of iGEM Marburg. Although his background is in nuclear and particle physics, his research interests shifted toward affordable biosensors for point-of-care cancer detection, which is how he ended up doing microbiology for iGEM.
Back in March when the iGEM season had just started, Maurice, a fellow iGEM Marburg team member, told me that he was exchanging emails with Margaretha Schwartz from Promega. Given my background as a physics student, Promega was not a household name for me at the time. “So, are you interested in automating a plasmid purification protocol?” asked Maurice. He told me that Promega was willing to supply the Wizard® MagneSil® Plasmid Purification System for this purpose; that was another name that added to my confusion.


This year, iGEM Marburg is aiming to establish a fast phototrophic organism as a synthetic biology chassis. For this goal we chose Synechococcus elongatus UTEX 2973, with a reported doubling time of 90 minutes. More specifically, we are creating an easy to use toolbox to empower rapid design testing, including genome engineering tools, self-replicating plasmid systems, natural competence and a Golden Gate-based part library. Our team chose to work on phototrophic organisms because we envision accelerating research in this particular field. (Note: Last year, Marburg’s iGEM project won the Grand Prize!)
Fast forward two months and I was beginning to understand that for Golden-Gate cloning or many other biological workflows, plasmid purification plays a crucial role. I shadowed Annsophie, a team member with more biology background, as she demonstrated a manual plasmid purification protocol. It was a long process (around four hours during our first try) and she told me that it is a common technique. Even after doing it several times and getting a bit faster, we still needed three hours, more or less. I thought it would be a relief if we could delegate this to our Opentrons OT-2 liquid handler, as it is quite mindless work.

Annsophie and I were then tasked to create an OT-2 protocol for plasmid purification. First we needed the equipment and expertise. We had a Skype session with Nans Bodet, a Field Support Scientist (FSS) from the automation department at Promega. Although he works with many different liquid handling instruments, Nans had not worked with an OT-2 before. He was very interested to see an OT-2 version of the protocol, as the OT-2 is much cheaper than other liquid handlers. He told us that a magnetic and shaker module for the OT-2 would be crucial for our protocol. He also suggested that if we wanted to increase our yield we would need an 8-channel pipette arm for our OT-2, which we did not have at the time. If successful, this project has the potential to serve the amateur market, including biohackers, enthusiasts, and students.
So up we went collecting the needed equipment. Nans recommended the D30-Elm from Qinstrument for our shaker. Since our team did not have the budget to purchase it (~€3000), I wrote their customer service to see whether they were willing to sponsor us. A member of their support team wrote us back and kindly helped us to secure a permanent loan for the device back in June.
Then I wrote to Opentron, who knew our team from last year, to talk about Marburg’s plan for plasmid purification. They were excited about the plan and helped us by providing the magnetic module. Moreover, our team won a grant large enough to purchase the 8-channel pipette arm. By August, the puzzle pieces were coming into place.
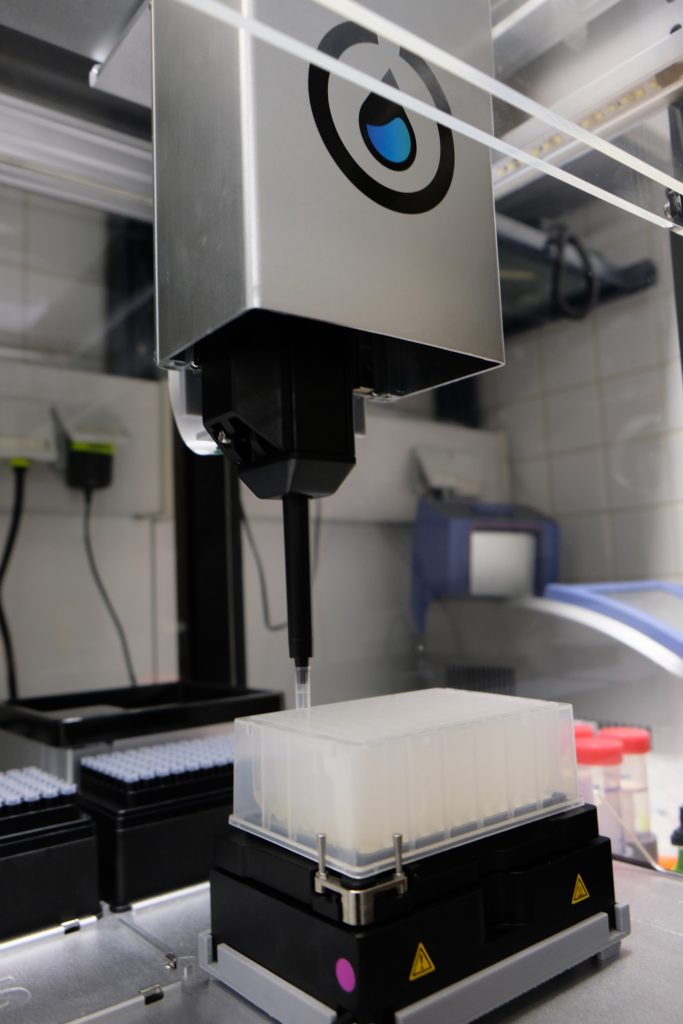
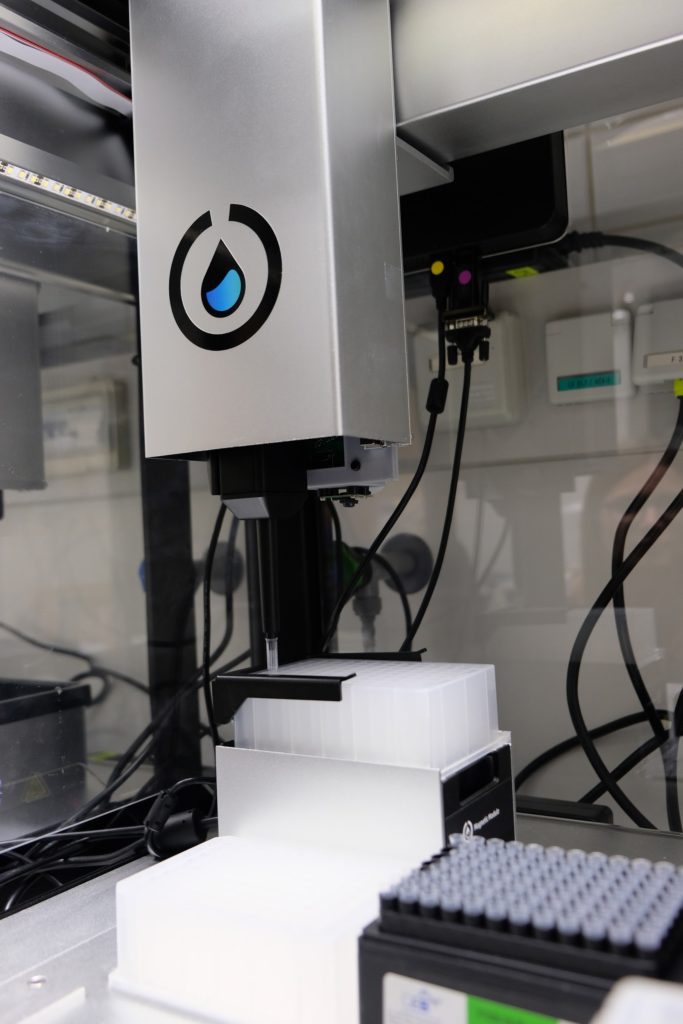


In the meantime, while waiting for all of the equipment to arrive at our lab, Annsophie and I began conceiving the protocol. She explained to me what would be needed to be done from a biological perspective and I tried to come up with the possible technical translation. Given her background in bioinformatics and mine in physics we covered much needed ground. Another team member, Cedric, helped by giving us feedback and designing a holder for the shaker to fit. The shaker was a bit bigger than the space normally occupied by modules in the OT-2 and needed stabilizing support.
Our 8-channel pipette arm arrived last, hence in August we tried doing 6 samples with a single-channel pipette arm. Since we did not want to waste any of our precious reagents, we did a proof-of-concept experiment using water. During this time, Opentron was rolling out a major update from their OT-2 3.9 to 4.0 firmware that included a lot of paradigm change.
This made defining labware difficult and we ended up defining our shaker module coordinates as a python dictionary importable via a .json file. So far this has worked perfectly. We also had many calibration problems with our OT-2 while trying to finish the protocol; thankfully Opentron customer service was patient with us. They told us how to calibrate the OT-2 directly via the terminal because we had many difficulties calibrating with the application.
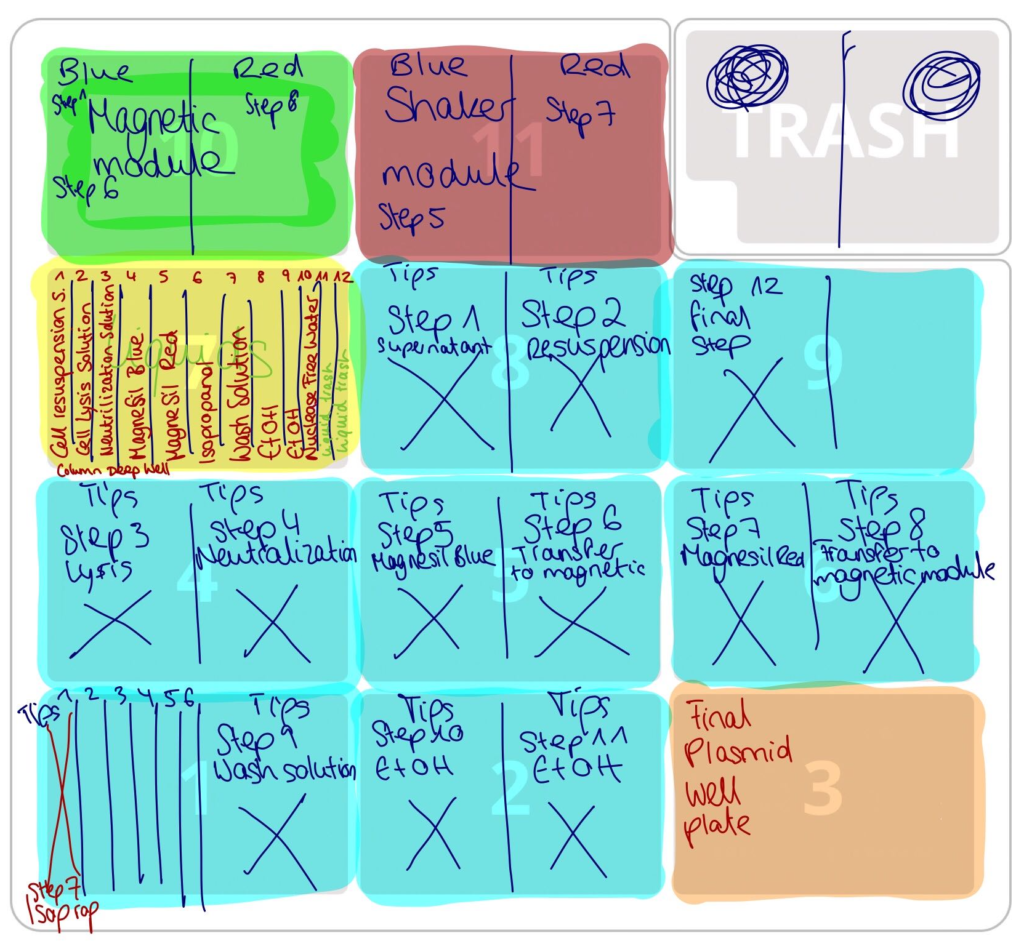
Now (as of the end of September) Annsophie and I have a working protocol for use with a single-channel pipette for up to 6 samples, as well as an 8-channel pipette for up to 32 samples. We have not tested the 8-channel pipette yet, but the single-channel protocol has successfully purified a sample, with virtually no intervention. While the yield that we got is still quite low (~ 60 ng/µl), we still have a lot of variables to play around with, especially incubation time for lysis.
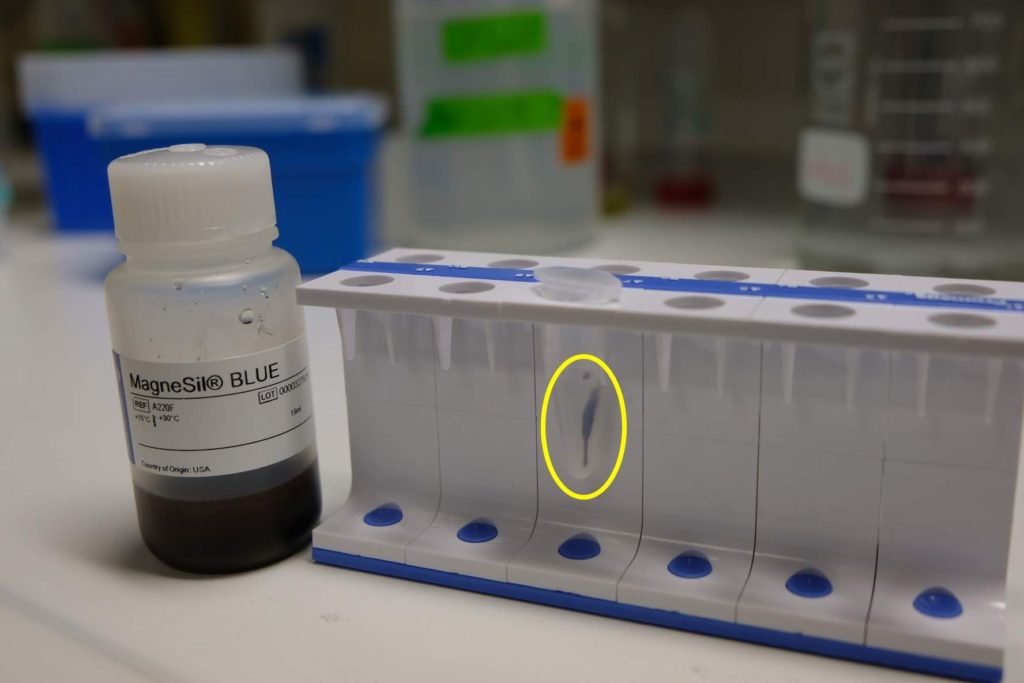
We noticed that during one trial we incubated our samples with lysis solution for too long (6 minutes), which resulted in an even lower yield. Another time incubation was too short (1 minute) and we noticed that this caused clumping of the MagneSil® beads, presumably because of excess protein and cell residues in the solution. This provides ample room for development and optimization as the iGEM deadline quickly approaches. We can’t wait to present the results of our project later this month at the 2019 iGEM Giant Jamboree.
Competing in iGEM? Learn more about how you can get support for your iGEM project at our website and check out our resources for iGEM teams, including Q&A webinars and tips for creating scientific posters.
Latest posts by Promega (see all)
- Accurate and On-Time: A Look Inside Promega Logistics - June 24, 2025
- Augmenting Human Capabilities with AI Tools - June 17, 2025
- Lead with Empathy: Supporting Caregivers in the Workplace - June 3, 2025
