Now that Promega is expanding its offerings of options for examining live-cell protein interactions or quantitation at endogenous protein expression levels, we in Technical Services are getting the question about which option is better. The answer is, as with many assays… it depends! First let’s talk about what are the NanoBiT and NanoBRET technologies, and then we will provide some similarities and differences to help you choose the assay that best suits your individual needs.
A Little Background on NanoBiT
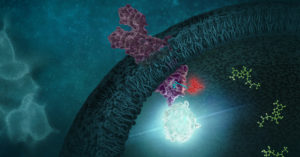
The NanoLuc® Binary Technology (NanoBiT®) is composed of two subunit bits that we call Large BiT (LgBiT; 18kDa) because it is bigger, and Small BiT (SmBiT; 1.3kDA) because it is little. These two subunits were individually optimized and have a very low affinity, reversible interaction that forms the bright NanoBiT® luminescent enzyme when in proximity to one another. When you tag your protein pair of interest, one with SmBiT and the other with LgBiT, the affinity of your proteins (if they interact) drives the interaction of the subunits and the protein-protein interaction (PPI) can be monitored in live cells using an aqueous, cell-permeable furimazine substrate. The new Bi-directional (BiBiT) vectors allow you to express both SmBiT and LgBiT fusions using a single vector for simpler stable cell line creation.
There is also the tiny HighBiT (HiBiT; 11 amino acid) subunit with a high-affinity for LgBiT, which is so small that you can use CRISPR/Cas9 genome editing to insert the tag to detect endogenous protein levels or stably insert HiBiT into a viral genome! This BiT is not good for protein interaction studies because of its high affinity. Instead, a HiBiT tag allows you to quantitate your protein of interest. You can detect extracellular proteins on live cells, proteins secreted into the medium, proteins in a cell lysate using a plate-based format, or a quickly detect tagged proteins after blotting to a membrane using HiBiT detection reagents that contain LgBiT. You can also detect intracellular proteins in live cells by expressing LgBiT in your system allowing quantitative live-cell kinetic measurements. Keep an eye out for more BiTs yet to come!
An Introduction to NanoBRET Technology
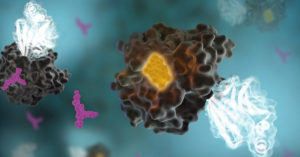
NanoBRET™ (Bioluminescence Resonance Energy Transfer) uses the bright and small NanoLuc luciferase as an energy donor for an acceptor fluorophore. In the case of protein-protein interaction studies, you express NanoLuc fused to one of your proteins of interest as the donor, and HaloTag® fused to the potential protein partner labeled and with the NanoBRET 618 fluorophore as the acceptor. To transfer resonance energy, the donor must be within 10nm of the acceptor and in the proper orientation making the technique useful for measuring protein interactions. The reduced overlap of these donor and acceptor emission spectra provide a better signal-to-noise ratio than other BRET methods. We offer a variety of prebuilt NanoBRET Protein Interaction Assays.
NanoBRET can also be used for target engagement (TE) assays to measure compound binding at select target proteins in live cells. Your protein of interest fused to NanoLuc is the energy donor and a cell-permeable fluorescent tracer attached to a test compound is the acceptor. Compounds that specifically engage the intracellular target protein-NanoLuc fusion will result in a decrease in BRET allowing real-time kinetic measurements. We have a wide variety of NanoBRET TE Intracellular Kinase Assays already optimized to measure kinase-ligand affinity in live cells!
Deciding Between BiT and BRET
Both the NanoBiT and NanoBRET technologies are great for measuring protein-protein interactions, so how do I choose for this application? The first thing to ask is whether your lab is familiar with resonance energy transfer assays and has the specific instrumentation available to detect this type of interaction. NanoBRET™ assays require an instrument capable of sequentially measuring dual-filtered luminescence and the appropriate filters. The ideal filter setup will include a band pass (BP) filter centered around 460nm to measure the donor signal (e.g., Emission 450nm/BP 80nm) and a long pass (LP) filter starting at around 600–610nm to measure the acceptor signal (e.g., Emission 610nm/LP). NanoBiT assays only require a standard instrument capable of measuring luminescence. With specialized imaging equipment, in vivo or in vitro imaging is also possible with both technologies. Here is a table of some similarities and differences to help you choose the assay that best suits the goals of your experiments.
How does NanoBiT® compare to NanoBRET™ for looking at protein-protein interactions (PPIs)?
| NanoBRET and NanoBiT Technologies | |
| Similarities | |
| Live cell kinetic assays possible | |
| Transfection based cloning | |
| Requires testing all 8 possible combinations of labeled protein pairs | |
| Stable cells require expression of two partners | |
| Bioluminescent imaging (BLI) possible with specialized imaging instruments | |
| Differences | |
| NanoBRET™ | NanoBiT® |
| Specialized filtered luminescence required | Standard luminescence required |
| The ratio of donor (NanoLuc fusion vector) to acceptor (HaloTag fusion vector) must be optimized | In general, a 1:1 ratio of SmBiT and LgBiT vectors is appropriate for PPI assays |
| Longer protocol due to multiple reagent additions | Shorter protocol due to single reagent addition |
| Ratiometric measurement that corrects for differences in cell number | Measures a gain or loss of signal so it is sensitive to differences in cell number |
| Utilizes proximity of protein pairs | Requires a physical interaction of subunits |
| Many ready-made assays | Fewer ready-made assays |
Whether you pick a BiT or BRET, we are happy to discuss these options in further detail and provide additional resources to help plan your project for success! You can reach the scientists in Technical Services by phone, chat, or email and we can even offer personalized seminars for your lab through the CoLABorate Training Program. We look forward to talking tech with you!
Latest posts by Joliene Lindholm (see all)
- A Day in the Life of a Technical Services Scientist - May 24, 2021
- Oh, The Ways You Can “Glo” - February 22, 2021
- Designing a Reporter Construct for Analyzing Gene Regulation - December 10, 2019