Bioluminescent reporter assays are an excellent choice for analyzing gene regulation because they provide higher sensitivity, wider dynamic range and better signal-to-background ratios compared to colorimetric or fluorescent assays. In a typical genetic reporter assay, cells are transfected with a vector that contains the sequence of interest cloned upstream of a reporter gene, and the reporter activity is used to determine how the target sequence influences gene expression under experimental conditions. A second control reporter encoded on the same or a different plasmid is an essential internal control. The secondary reporter is used to normalize the data and compensate for variability caused by differences in cell number, lysis efficiency, cell viability, transfection efficiency, temperature, and measurement time.
For genetic reporter assays, using a secondary control vector with a weak promoter like PGK or TK to ensures that the control does not interfere with activation of your primary reporter vector. Transfection of high amounts of the control plasmid or putting the control reporter under control of a strong promoter like CMV or SV40 often leads to transcriptional squelching or other interference with the experimental promoter (i.e., trans effects). Reporter assays can also be used to quantitatively evaluate microRNA activity by inserting miRNA target sites downstream or 3´ of the reporter gene. For example, the pmirGLO Dual-Luciferase miRNA Target Expression Vector is based on dual-luciferase technology, with firefly luciferase as the primary reporter to monitor mRNA regulation and Renilla luciferase as a control reporter for normalization.
Here in Technical Services we often talk with researchers who are just starting their project and looking for advice on designing their genetic reporter vector. They have questions like:
- How much of the upstream promoter region should be included in the vector?
- How many copies of a response element will be needed to provide a good response?
- Does the location of the element or surrounding sequence alter gene regulation?
Optimize the Construct for Your Experiment
Biology is complex, so there is no single correct answer for these questions, but there are many resources and guidelines to help you optimize the sequence that will best suit your experiment. One advantage of genetic reporter assays is that you can control which elements are examined.
- You might include the entire proximal promoter (including ~1kb upstream of the promoter), a specific promoter subsection or as little as a single response element.
- When generating a transcriptional fusion or using a vector with no transcription start site, you might include the +1 transcription start site.
- The reporter might include the 5´ untranslated region (UTR) if you want to understand how this sequence may affect promoter activity, but keep in mind that UTR sequences also can affect post-transcriptional regulation.
- The reporter might include an intron and all or part of one or both flanking exons to study RNA splicing or characterize regulatory elements contained within the intron (but if the exons include coding sequence, be sure to clone the reporter gene in frame).
- The reporter might contain the 3´ UTR to focus on post-transcriptional regulation through miRNA effects.
- You might clone any combination of sequences from your gene of interest to look at the integration of regulatory pathways; reporter assays allow this flexibility and refined experimental design.
Step One: Stand on the Shoulders of Giants

The first guideline for deciding which sequence to incorporate is to search the literature to find out what is known about your gene of interest. Often, the minimal required upstream region has been identified in many model organisms, and you can use that to focus your efforts. Identifying a positive control treatment known to alter transcription is a valuable tool for examining gene regulation, even if the mechanism is unknown. The National Center for Biotechnology Information (NCBI) is a good place to start to look up your gene and associated publications. You may even be able to find a vector that has worked for other researchers.
If nothing is known about the promoter of your gene of interest, a crude but tried-and-true method is to just use 1–3kb upstream of your gene. You can then start reducing the size of the insert sequence and trying different segments to narrow down the important elements. If there is a known response element, but one copy doesn’t give a good signal, you can try adding multiple copies of the element, which may enhance the signal. There are many databases, tools and resources available on the topic of gene regulation. The Encyclopedia of DNA Elements (ENCODE) projects provide publicly available data for human, mouse, nematode and fruit fly genomes. There are many other options, and new tools are always being developed. Reach out to experts in your field to see what new options are available.
Step Two: Choose the Vector Backbone
Once you have decided what sequence to include, the next step is deciding which reporter vector backbone to use for your experiments. We recommend the brightest and most sensitive NanoLuc® luciferase for the primary reporter gene.
Typically, researchers will choose between a promoterless or a minimal promoter vector. Both options with no insert (empty vectors) can provide the desired low basal level of background reporter signal, but the signal will vary depending on the cell line and transfection conditions. If your sequence of interest is expected to just enhance transcription, you may need to use a minimal promoter vector to achieve a basal signal above background. If your sequence of interest is a promoter region and has all the necessary components for basal transcription, the promoterless vector will likely be suitable.
Step 3: Consider Additional Vector Features
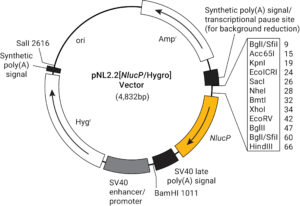
Next, you can choose between other options like a reporter modification. For analyzing gene regulation, we always recommend using a reporter protein modified to have a short half-life. This ensures that the signal will not build up as the protein accumulates over time. Using a reporter with a short protein half-life more closely ties protein expression to changes in transcriptional activity and increases signal-to background ratios. The destabilized versions of the luciferase reporters (luc2P, luc2CP, hRlucP, hRlucCP and NlucP) enhance reporter response dynamics, reduce assay time and minimize secondary effects that may arise from the prolonged incubation of cells with screening compounds.
You can also choose to use a vector with an antibiotic resistance selectable marker, which can be used to select for cells that have stably incorporated the vector by treating your cells with an antibiotic that will kill the cells without the vector. Use whichever selectable marker (hygromycin, puromycin or neomycin) is most convenient for your experiment and cell type. Even if you don’t plan to make stable cells right now, it never hurts to have it in your vector in case you decide to make stable cells in the future.
Step 4: Analyze the Data
Researchers often want to know what a good background or basal signal value should be, but instruments calculate relative light units (RLUs) by different methods and since it is a relative value, the raw signal cannot be compared across instruments, even of the same model. A more useful comparison is the RLU of cells transfected with your vectors versus untransfected cells.
A solid white plate should be used for your reading your samples because it will provide the best signal and lowest crosstalk (light leaking into adjacent wells). The dynamic range and sensitivity of an instrument is a key factor when measuring the extremes of a very low or a very high signal. A sensitive instrument will have a clear distinction between the blank noise and the sample signal. As with any luminescent assay, we do not recommend comparing the raw RLU values. Instead, we compare the ratio of primary reporter to control reporter for genetic reporter assays.
When you have your sequences and the reporter vector that will best suit your needs, you will be ready to start cloning (that is a whole other topic). As always, please contact Technical Services if you have any questions before you get started or at any time as you move forward with your project.
Related Posts
Latest posts by Joliene Lindholm (see all)
- A Day in the Life of a Technical Services Scientist - May 24, 2021
- Oh, The Ways You Can “Glo” - February 22, 2021
- Designing a Reporter Construct for Analyzing Gene Regulation - December 10, 2019
