MicroRNAs (miRNAs) are small, non-coding RNAs that play a role in regulating cancer by acting as both tumor suppressors and oncogenes. Ranging in size from 18–25 nucleotides, miRNAs function in feedback mechanisms to regulate many cellular processes including cell proliferation, apoptosis, cell signaling and tumorigenesis (1).
Not surprisingly, dysregulation of miRNA expression can have serious repercussions. For example, miRNAs are dysregulated in almost all human cancers (1). Because of the potential to influence cancer growth and development, there is growing interest in miRNA profiling to identify possible biomarkers for cancer diagnosis or prognosis, as well as potential therapeutic targets (1).
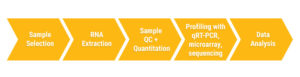
Growing interest in miRNAs as both biomarkers of disease and therapeutic targets drives the need for fast and effective methods for miRNA profiling. Profiling miRNA targets follows a relatively simple workflow: sample selection, RNA extraction, RNA QC and quantitation, RNA profiling and data analysis (2,3). So what happens at each step?
Sample Selection
Depending upon your area of focus, you could be extracting RNA from a variety of sample types. High-quality miRNA can be extracted from both cells and tissue relatively easily. Because miRNAs are surprisingly stable in FFPE tissues, these tissues also yield usable quantities of miRNA. The cell heterogeneity of your tissue is also an important consideration because many miRNAs are tissue-specific. For this reason, using microdissection methods or other cell-targeting approaches are advisable. Isolating miRNA from blood plasma is challenging in part because of the endogenous RNase activity. The white blood cells and red blood cell hemolysis can also adversely affect the quality and quantity of miRNA extracted.
RNA Extraction
The principles for extracting miRNA is generally the same as those for extracting RNA with the exception that they are modified to capture more of the small RNA fraction. For some sample types, miRNA extraction methods may require optimization. For example, in their native state, endogenous blood plasma miRNAs are protected from RNase; however, extracted miRNAs that are spiked into plasma degrade quickly. For this reason, it is important that any technique for miRNA extraction form blood plasma quickly and completely inactivate the RNase activity.
RNA QC and Quantitation
Evaluating the quality of your extracted RNA is important for both the quality and reproducibility of your miRNA profiling results. Because most profiling methods use total RNA, it is not usually necessary to quantitate the amount of miRNA present in your extracted RNA (2).
RNA Profiling Methods
qRT-PCR
Today, quantitative RT-PCT (qRT-PCR) is considered the gold standard because it is the best at absolute miRNA quantification and offers the greatest sensitivity and dynamic range. In this technique, miRNAs are reverse transcribed into cDNA by first adding a poly(A) tail using E. coli poly(A) polymerase. The small size of miRNAs won’t allow traditional reverse transcriptase primers to bind. Instead, a reverse transcription primer binding site needs to be created using a special stem looped primer. Following reverse transcription, amplification can be performed using traditional real-time PCR methods that measure the accumulation of the amplified product in real-time (2).
Microarrays
One of the first methods used for parallel analysis of large numbers of miRNAs is microarray analysis. MiRNAs are tagged using fluorophore-labeled nucleotides and then hybridized to capture probes located or “arrayed” on slides or beads. After hybridization, fluorescent detection reveals what miRNAs are present. Microarrays can’t be used to quantitate the amount of a specific miRNA and are best used for comparing the relative amount of miRNA present between two states (e.g., normal and diseased tissue).
RNASeq
Based on next-generation sequencing (NGS) technology, RNAseq involves preparing a cDNA library using the RNA sample and then performing “massively parallel” sequencing of the millions of cDNA molecules all at one time. This method can identify known and new miRNA sequences in a high-throughput manner and provides a relative quantification of each miRNA sequence (number of an individual miRNA reads compared to the total reads in a sample). RNASeq methods can identify many possible new miRNA species, but they may not all be actual miRNAs, so confirmation is required (2).
Data Analysis
Analysis of miRNA profiling data is divided into several steps: processing, quality assessment, normalization and differential expression calculations. The best approach to these steps is dependent on the miRNA profiling platform and the goal of the experiment (2).
The growing interest in the role miRNA plays in disease development means that methods available for each step of this workflow will continue to develop and improve.
References
- Peng, Y. and Croce, C. (2016) The role of MicroRNA in human cancer. Signal Transduction and Target Therapy. Jan 28;1:15004.
- Tewari, M et al. (2015) MicroRNA profiling: approaches and considerations. Nat Rev Genet. 13, 358–369.
- Liu, F. et al. (2016) The complexity of microRNAs in human cancer. J. Rad. Research. 57, i106–i11.
Kelly Grooms
Latest posts by Kelly Grooms (see all)
- Growing Our Understanding of Triple-Negative Breast Cancer in Sub-Saharan Africa: Why Comprehensive Population Data Matters - June 5, 2025
- Measles and Immunosuppression—When Getting Well Means You Can Still Get Sick - April 17, 2025
- IC50, EC50 and Kd: What is the Difference and Why Do They matter? - March 6, 2025
