The SARS-CoV-2 nucleocapsid protein accounts for the largest proportion of viral structural proteins and is the most abundant protein in infected cells. Nucleocapsid proteins have the job of “packaging” the viral nucleic acid (in this case, RNA). Viral nucleocapsid proteins can also enter the host nucleus and interact with a variety of host proteins to interfere with critical processes of the host cell, including protein degradation. Here we review a study that used an in vitro protein degradation assay to investigate the interaction of the SARS-CoV-2 nucleocapsid protein and the proteasome activator PA28γ.
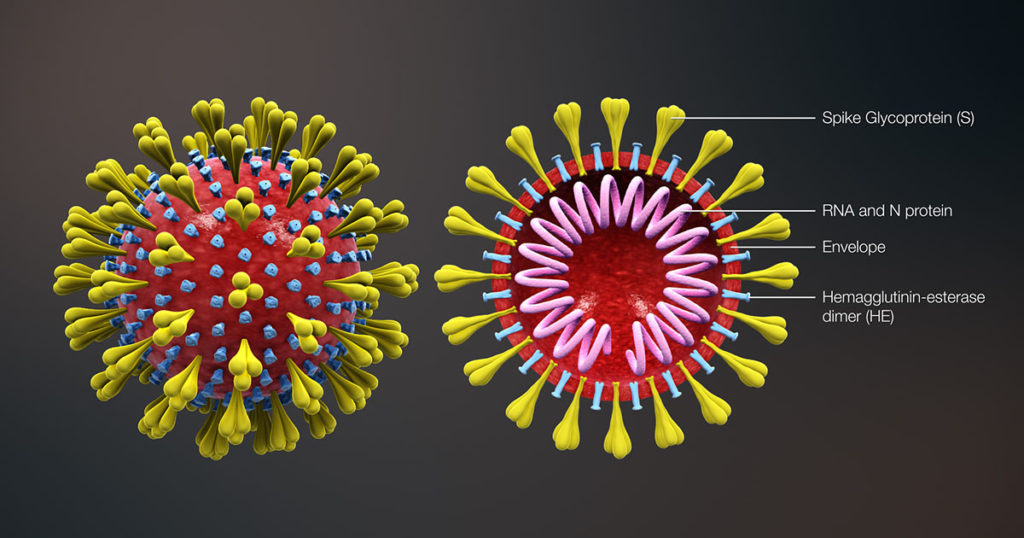
In SARS-CoV-2 infections, the nucleocapsid protein is critical for infection, replication and packaging. The SARS-CoV-2 nucleocapsid protein is not only localized in the cytosol of the host cell but also is translocated into the nucleus. There, it interacts with various cellular proteins that modulate cellular functions, such as the degradation of unneeded or damaged proteins by proteolysis. Researchers have proposed that the protein degradation system plays an important part in coronavirus infection (1).
Several studies have shown that the proteasome is important for viral infection. For instance, hepatitis C virus (HCV) core proteins can be degraded through the PA28γ-20S proteasome pathway. Stably maintaining HCV core proteins in the nucleus of infected cells is important to pathogenesis of HCV (2). In a separate study, PA28γ acted as a co-repressor of Human T-cell Leukemia Virus (HTLV-1) p30 protein to suppress viral replication and control latency (3).
The SARS-CoV-2 nucleocapsid protein may also interact with the proteasome activator PA28γ, which is localized in the nucleus. PA28γ could be critical for degrading the nucleocapsid protein in the nucleus as part of the 20S proteasome, which acts to degrade proteins in a ubiquitin-independent manner. Regulating the abundance of the critical nucleocapsid protein during infection via protein degradation could play a major role in SARS-CoV-2 pathogenesis.
Learn more about targeted protein degradation for drug discovery: https://www.promega.com/products/small-molecule-drug-discovery/protein-degradation-drug-discovery/
In a recent publication, using both in vivo and in vitro experiments Zhang and colleagues present data suggesting that PA28γ could mediate the degradation of SARS-CoV-2 nucleocapsid protein. (4)
A protein degradation in vitro assay was key to establish this correlation (5). The SARS-CoV-2 nucleocapsid protein was created by in vitro translation using a TNT® in vitro translation system. Purified PA28γ and 20S proteasome proteins were added to the synthesized protein. SARS-CoV-2 nucleocapsid exhibits down-regulation in the presence of PA28γ and the 20S proteasome, while PA28γ or 20S proteasome alone serves no function in the degradation of the SARS-CoV-2 nucleocapsid protein.
According to the Johns Hopkins coronavirus resource center, as of December 9, the COVID-19 pandemic has claimed the lives of more than 1.5 million people worldwide and left many more struggling with long-term effects of the infection. Understanding the biology of the SARS-CoV-2 virus is critical to developing multiple strategies for treatment and control.
Learn more about our products to support Viral Research, Vaccine and Therapeutic Development.
- Raaben, M. et al. (2010) The ubiquitin-proteasome system plays an important role during various stages of the coronavirus infection cycle. J. Virol. 84, 7869–79.
- Moriishi, K. et al. (2003) Proteasome activator PA28gamma-dependent nuclear retention and degradation of hepatitis C virus core protein J. Virol. 77, 10237–49.
- Ko, N-L. et al. (2013) PA28γ is a novel corepressor of HTLV-1 replication and controls viral latency. Blood 121, 791–800.
- Zhang, H et. al. (2020) Proteasome activator PA28γ-dependent degradation of coronavirus disease (COVID-19) nucleocapsid protein: Biochem. Biophysical Res. Comm. 529, 251e256
- Song, P. et. al. (2016) Parkin Modulates Endosomal Organization and Function of the Endo-Lysosomal Pathway 36, 2425-2437
