Today’s blog is written by guest blogger Kristin Huwiler from our Cellular Analysis and Proteomics Group.
Two research collaborations, one in Europe and a second in the US, have just published in Nature Chemical Biology (1,2) on the identification of BET inhibitors (bi-BETs) that bind via a bivalent mechanism to both bromodomains of BRD4. These bivalent chemical inhibitors exhibit high cellular potency and affinity relative to their monovalent predecessors. By developing high-affinity ligands that engage both bromodomains simultaneously within BRD4, the authors illustrate a concept that may be applicable in the development of selective, potent ligands for other multi-domain proteins. Here we review the work presented in the Waring et al. paper using the Promega NanoBRET™ Technologies to characterize the mechanism of action of their bivalent probe.
The bromodomain and extraterminal (BET) sub-family are some of the most studied bromodomain-containing proteins (3). The BET subfamily of proteins contain two separate bromodomains. BRD4 is one well studied member of the BET sub-family. Several small molecule inhibitors that target BRD4 have been developed as potential therapeutics for various cancers with promising initial studies, but to date are all monovalent, binding each bromodomain of the BET family members separately (2).
Proving the Bivalent Mechanism of Action
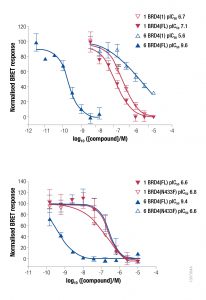
Waring et al. fully characterized these chemical probes using biochemical, biophysical and cellular assays. Several of the cellular assays used Promega technologies to provide evidence that the bivalent binding mechanism occurred in cells as did increased cellular affinity and residence time. The technologies and assays used included: NanoBRET™ Protein:Protein Interaction Assays, NanoBRET BioSensors, and NanoBRET Target Engagement Assays. Waring et al. tested multiple BRD4 constructs including; full-length, wild type BRD4 or point mutations within each respective bromodomain known to cause loss of binding to acetylated lysine, the tandem bromodomains with and without mutations, and the segregated bromodomains.
NanoBRET™ PPI Experiments
NanoBRET Protein-Protein Interaction (PPI) technology demonstrates the cellular potency of these compounds at disrupting the interaction between BRD4 and histone H3. Bivalent compound #6 in the Waring et al. paper showed increased cellular potency, by several orders of magnitude, for disrupting chromatin occupancy relative three other conditions: the isolated bromodomain 1, a point mutant, and treatment with monovalent compound #1.
NanoBRET™ BioSensor Experiments
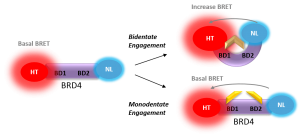
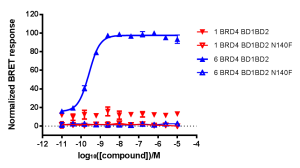
NanoBRET Biosensor demonstrated that BRD4 underwent a conformational change in cells when it bound bivalent compound #6, but not with monvalent compound #1. An increase in BRET signal was observed with increasing concentrations of compound #6 in a sensor containing both wild-type domains of BRD4. No change in signal was observed with increasing concentrations of monovalent binding compound #1
NanoBRET Target Engagement Experiments
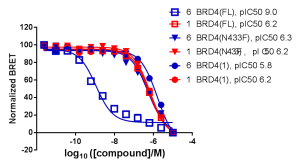
NanoBRET Target Engagement (TE) demonstrated that the intracellular affinity of full-length, wild type BRD4 for bivalent compound #6 was several orders of magnitude greater than that of monovalent compound #1. The isolated bromodomain 1 of BRD4 showed similar affinities for compounds #1 and #6.
Additionally, NanoBRET TE demonstrated that compound #6 had a longer intracellular residence time, compared to compound #1, for full-length BRD4. No difference in intracellular residence time for compound #6 relative to compound #1 was observed for the isolated bromodomain 1 of BRD4 or full-length BRD4 mutant N433F (data shown in Waring et al. 2016).
Summary
These and other experiments detailed in the Waring et al. paper describe the biochemical and cellular mechanism of action of a novel class of BRD inhibitor (3). The strong inter-assay correlation observed among the NanoBRET intracellular assays supports the proposed bivalent molecular mechanism of action of this novel chemical probe. This increased understanding of mechanism of action of a small molecule and the ability to “see” how it acts inside the cell will enable more informed and better therapeutic design.
Literature Cited
- Waring, et al. (2016) Potent and Selective Bivalent Inhibitors of BET Bromodomains. Nat. Chem. Bio. (Advanced Publication Online; DOI: 10.1038/nchembio.2210)
- Tanaka, M., et al. (2016) Design and Characterization of Bivalent BET Inhibitors. Nat. Chem. Bio. (Advanced Publicaiton Online: DOI: 10.1038/nchembio.2210)
- Prinjha, R.K. et al. (2012) Place your BETs: The Therapeutic Potential of Bromodomains. Trends Pharm Sci. 33: 146.
Updated April 4, 2024 to correct broken link.