 GPCRs
GPCRs
G protein-coupled receptors (GPCRs) are a large family of receptors that traverse the cell membrane seven times. Functionally, GPCRs are extremely diverse, yet they contain highly conserved structural regions. GPCRs respond to a variety of signals, from small molecules to peptides and large proteins. Many GPCRs are involved in disease pathways and, not surprisingly, they present attractive targets for both small-molecule and biologic drugs.
In response to a signal, GPCRs undergo a conformational change, triggering an interaction with a G protein—a specialized protein that binds GDP in its inactive state or GTP when activated. Typically, the GPCR exchanges the G protein-bound GDP molecule for a GTP molecule, causing the activated G protein to dissociate into two subunits that remain anchored to the cell membrane. These subunits relay the signal to various other proteins that interact with or produce second-messenger molecules. Activation of a single G protein can result, ultimately, in the generation of thousands of second messengers.
Given the complexity of GPCR signaling pathways and their importance to human health, a considerable amount of research has been devoted to GPCR interactions, both with specific ligands and G proteins.
BRET Who?
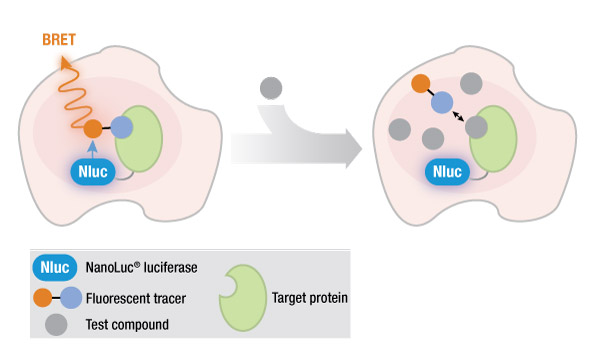
One method of studying protein interactions is known as bioluminescence resonance energy transfer (BRET), developed in 1999. Like its older cousin, fluorescence resonance energy transfer (FRET), BRET involves the nonradiative transfer of energy from a donor molecule to a fluorescent acceptor molecule that is in close proximity. The donor in BRET is luminescent (the signal is generated by the reaction of a luciferase enzyme with its substrate), while the donor in FRET is a fluorescent molecule with an excitation wavelength sufficiently different from the emission wavelength of the acceptor. This requirement for a difference between wavelengths is one of several disadvantages of FRET, compared to BRET.
However, the original BRET method suffered from a disadvantage when studying endogenous protein interactions in live cells: The BRET signal is relatively dim, due to the low light output of a standard luciferase reaction.
An effective solution to this problem was the introduction of a novel, engineered NanoLuc® luciferase that generates an exceptionally bright signal, making it possible to study GPCR-ligand binding in live cells using a NanoBRET™ assay. The small size of NanoLuc® luciferase (19 kDa) enables researchers to fuse the NanoLuc® protein to the protein of interest, with minimal effects on protein binding.
Mighty Nano
In a typical NanoBRET™ assay to study GPCR interactions, the NanoLuc® fusion protein is expressed within the cell, either transiently or stably, from a suitable vector. However, the high levels of protein expression from such a fusion construct may not accurately represent the true biology of an endogenously expressed GPCR. In 2017, White et al. published a novel method for fusing NanoLuc® luciferase(Nluc) to an endogenous GPCR gene using CRISPR/Cas9 gene editing technology (1). Their work used NanoBRET™ assays to study beta-arrestin recruitment by GPCRs, as well as GPCR internalization and trafficking. They reported that their method:
…overcomes the need for ectopic expression of the donor and therefore represents an advance in the current BRET proximity assays.
In the 2017 study (1), the authors generated C-terminal Nluc fusions for the proteins of interest. Recently, they extended their study (2) to demonstrate the successful generation of N-terminal Nluc fusions, using CRISPR/Cas9, in frame with the endogenous ADORA2B locus in HEK293 cells. This gene encodes the adenosine A2B receptor, a GPCR that plays a critical role in several diseases, including cancer, renal disease and diabetes.
Using NanoBRET™ assays to observe the binding of fluorescent ligand to the Nluc/A2B fusion receptors, the researchers demonstrated that:
…while these observations from gene-edited receptors are largely comparable to those from Nluc-tagged receptors that are over-expressed, they provide more consistent results when at similar levels of luminescence.
They also determined the pharmacological parameters of ligand binding for a nonspecific fluorescent adenosine receptor antagonist, XAC-X-BY630. Their results demonstrate the advantages of combining NanoBRET™ assays with gene-editing experiments to study Nluc fusion proteins under endogenous expression conditions, and should be broadly applicable to the study of other physiologically relevant membrane proteins.
For more information on using CRISPR/Cas9 technology to study protein interactions in live cells, see this webinar by Dr. Chris Eggers.
References
- White, C.W. et al. (2017) Using nanoBRET and CRISPR/Cas9 to monitor proximity to a genome-edited protein in real-time. Sci. Rep. 7, 3187.
- White, C.W. et al. (2019) NanoBRET ligand binding at a GPCR under endogenous promotion facilitated by CRISPR/Cas9 genome editing. Signal. 54, 27.
Latest posts by Ken Doyle (see all)
- CRISPR/Cas9 Knock-In Tagging: Simplifying the Study of Endogenous Biology - November 18, 2025
- Will Artificial Intelligence (AI) Transform the Future of Life Science Research? - February 1, 2024
- RAF Inhibitors: Quantifying Drug-Target Occupancy at Active RAS-RAF Complexes in Live Cells - September 5, 2023
