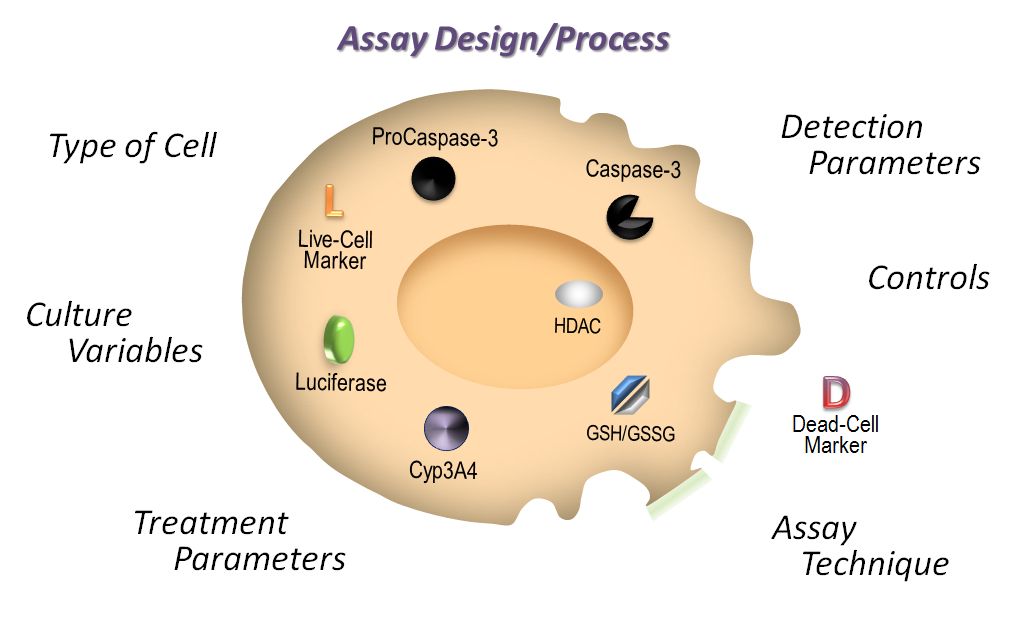
Welcome to the third installment of our series on cell-based assays; in this post we talk about treatment parameters for cell-based assays. Designed for the newbie to the world of cell-based assays, we have covered the topics of choosing your cell type and basic cell culture tips in the previous posts. In this post, we will discuss how decisions about test compound treatment: how much and how long can affect assay results and interpretation.
Drug Dose
The drug dose affects the timing and synchrony of the cellular response, and it can even determine the type of response. For instance, some drugs that induce apoptosis at low doses may be acutely cytotoxic at higher doses.
Timing
The optimal timing of treatment will be affected by the type of cell, the test compound, dosage and endpoint of the assay. For instance, if your assay measures caspase activity, you may need to look at earlier time points, because caspase activation is an early event in apoptosis. At later time points (24 to 48 hours) the cells may be dying through mechanisms of secondary necrosis, and assays for caspase activity would give falsely low results.
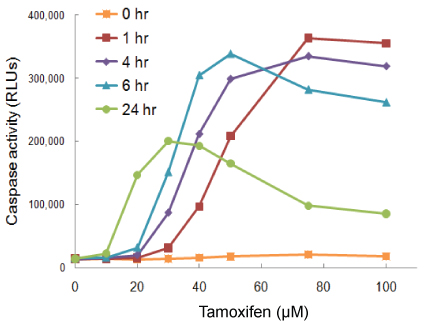
In this example cells were treated with tamoxifen and timing between treatment and the Caspase-Glo® 3/7 Assay was varied. At 1 hour, the maximum response is seen with the highest dosage of drug. At 24 hours, maximum responsiveness is seen with lower dosages. Because caspase is a transient analyte, at 24 hours in the assay, at higher doses most of the cells have likely already passed through the apoptotic pathway and gone on to secondary necrosis…so, you get a falsely low signal.
This experiment illustrates the importance of performing drug response curves and timing experiments. However, multiplexing assays with different endpoints can also help alleviate concerns about the effects of dosage and timing on your results.
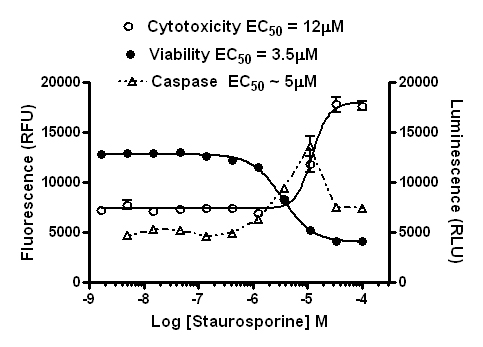
In this case, you see that at higher doses, at the time point the caspase assay was read, there was significant induction of apoptosis peaking at ~10µM, but at higher doses it isn’t clear apoptosis was induced from the caspase assay alone. Could primary necrosis be at work here?
However, based on the data from the other assays, you can see that most of the cells died off shortly after the caspase signal peaked. This would explain why the caspase signal diminished (the cells were dead). By comparing the different components of the multiplex in a graph like this you can determine that caspase activity was strongly induced by treatment, and apoptosis was the mechanism of cell death.
You can even normalize the data sets from the multiplexed assays. Cells undergoing apoptosis are still alive. When you normalize caspase activity to viability, you see that the live cells show a strong induction of apoptosis.
These data illustrate how multiplexing helps you avoid misinterpretation of data. If you miss the right combination of assay parameters, you essentially expand the “sweet spot” of your assay.
So in addition to doing the appropriate control experiments such as dose response curves and time course treatments to fully characterize your experimental system, multiplexing assays with a variety of endpoints allows you to compare multiple parameters from the same sample, giving you more information from your assay.
Articles:
Multiplexing Cell-Based Assays: Get More Biologically Relevant Data
Choosing the Right Cell-Based Assay for Your Research
Michele Arduengo
Latest posts by Michele Arduengo (see all)
- IL-6/STAT3-Regulated Long Non-Coding RNA Is Involved in Colorectal Cancer Progression - July 1, 2025
- Using Dual-Luciferase Assays to Identify the Role of Non-Coding RNAs in Disease - May 2, 2025
- An Unexpected Role for RNA Methylation in Mitosis Leads to New Understanding of Neurodevelopmental Disorders - March 27, 2025