The path to drug development is strewn with obstacles: Identifying targets; configuring assays to help identify targets or drugs; uncovering the right compound to affect the selected target without off-target effects and screening multiple compounds to eliminate or identify potential drugs. Without the right tools, compounds or target, identifying potential disease therapies becomes nearly impossible.
When it comes to a drug target for cancer, the Ras protein family is at the top of the list because the proteins are expressed ubiquitiously and found mutated in many types of cancer. Because Ras proteins are involved in transducing signals from the surface of cells, many of the resulting mutations produce an activated Ras, inducing uncontrolled expression of the genes that Ras controls. Ras proteins are small GTPases (20–25kDa) that comprise a larger superfamily of proteins divided into five subfamilies: Ras, Rho, Rab, Arf, and Ran. These proteins control diverse cellular activities, including cellular differentiation, proliferation, cell division, nuclear import and export, and vesicle transport. GTPases are guanosine-nucleotide-binding proteins with affinity for GDP or GTP and are able to hydrolyze GTP. When bound to GTP, GTPases are active (turned on) and interact with downstream proteins in the signaling cascade. When GTPases are bound to GDP, the proteins are inactivated (turned off) and no longer transduce signals.
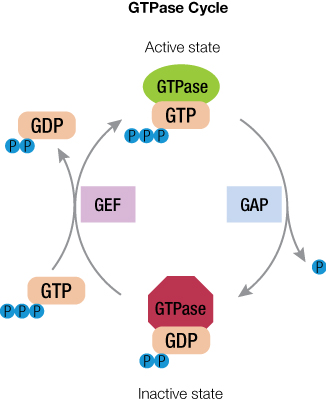
This GTPase on-off state is regulated by guanine nucleotide exchange factors (GEFs) and GTPase activating proteins (GAPs). When the inactive GTPase bound to GDP is stimulated, GEF activates the GTPase to eject GDP and then binds GTP because GTP concentration is 10X higher than GDP inside the cell. The activated GTPase bound to GTP then acts in the cellular signal cascade. In general, GTP hydrolysis by GTPase is a slow process, but in the presence of GAPs, hydrolysis is accelerated, resulting in GTPase bound to GDP. This inactivates the GTPase until the next encounter with GEF. Thus, GTPase has a cycle of on and off states, depending on cellular signaling. And each of these players (GTPase, GAP and GEF) are potential targets for drug development.
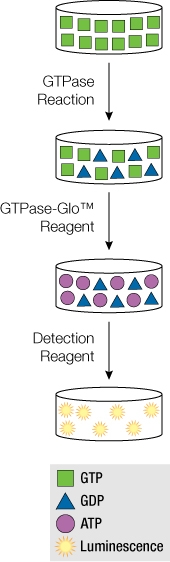
While Ras and the other small GTPases are important proteins to study, tools for studying GTPases and their regulatory proteins GEFs and GAPs have lacked convenient assays. Our proven luminescence-based technology has been used to develop the GTPase-Glo™ Assay. This assay can be used for studying the activity of GTPases, GAP-assisted GTPases, GAPs and GEFs. Based on the principle of GTP hydrolysis of GTP results in GDP and Pi, the GTP remaining after enzyme activity is measured and quantitated in a luminescent reaction. Because more active enzymes use more GTP, this results in a lower luminescent signal. Thus the light output from the GTPase-Glo™ Assay shows an inverse correlation to enzyme activity.
Using purified proteins and a reaction buffer specific for each GTPase reaction (e.g., low magnesium concentration eliminates the need for GEFs in the GTPase reaction), the GTPase reaction starts once GTP is added. When the GTPase reaction is complete, add an equal volume of GTPase-Glo™ Reagent, mix and measure luminescence. The sensitive assay can be used for high-throughput processing, making that search for the potential drug quicker.
The demonstration of the GTPase-Glo™ Assay was published in Assay and Drug Development Technologies. Read the peer-reviewed article here.
Sara Klink
Latest posts by Sara Klink (see all)
- A One-Two Punch to Knock Out HIV - September 28, 2021
- Toxicity Studies in Organoid Models: Developing an Alternative to Animal Testing - June 10, 2021
- Herd Immunity: What the Flock Are You Talking About? - May 10, 2021