 Cancer. It has been the nemesis of medical science for decades. We declare war on it, we wax philosophical about finding a cure for it. We talk about it as if it were a single enemy, but it isn’t—Cancer is not a disease, it is hundreds of diseases. These diseases manifest in every region of the human body. Many cancers, if diagnosed early, can be treated successfully. Unfortunately, there are many forms of cancers that have no external signs and very few symptoms at the early stages. Such is the enemy we face: we know what it is, we know we need to kill it early and in many cases we know how to do it, what we don’t know is how to catch it early.
Cancer. It has been the nemesis of medical science for decades. We declare war on it, we wax philosophical about finding a cure for it. We talk about it as if it were a single enemy, but it isn’t—Cancer is not a disease, it is hundreds of diseases. These diseases manifest in every region of the human body. Many cancers, if diagnosed early, can be treated successfully. Unfortunately, there are many forms of cancers that have no external signs and very few symptoms at the early stages. Such is the enemy we face: we know what it is, we know we need to kill it early and in many cases we know how to do it, what we don’t know is how to catch it early.
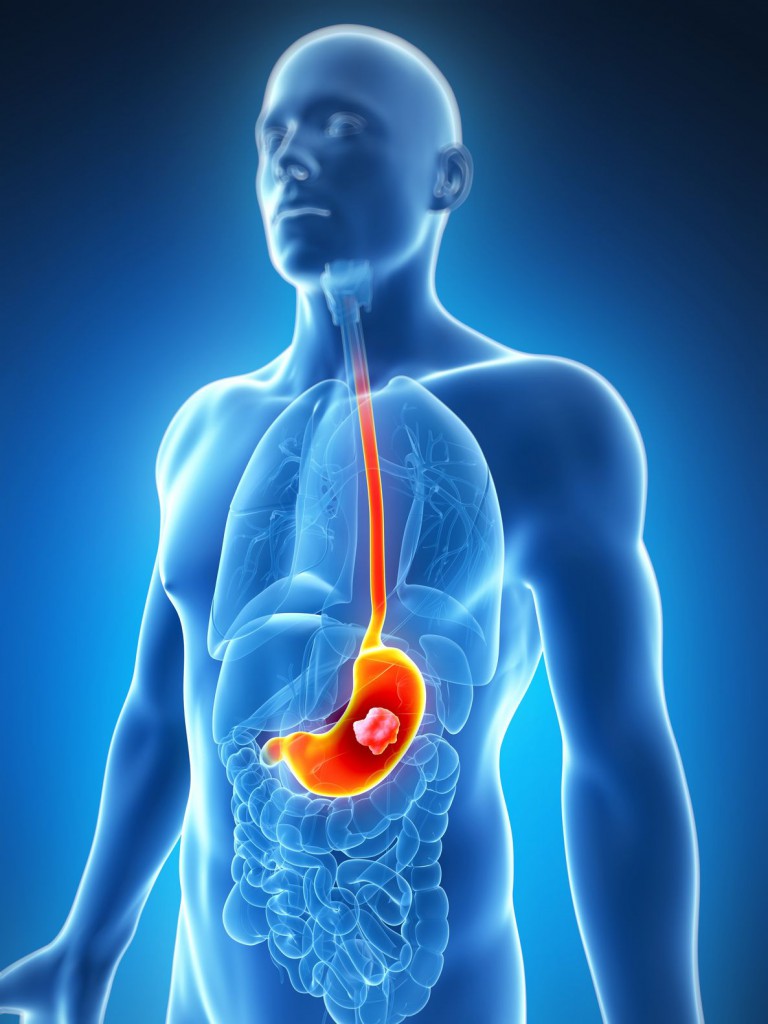
Gastric cancer kills approximately 745,000 people a year worldwide, making it the third most common cause of cancer-related deaths (1). It has such a high mortality rate because usually it is not detected until the disease has progressed to the later stages (IIIA–IV; 2). When detected this late, the 5-year survival rate ranges from 7–27%, with the median survival being less than 12 months (2). In contrast, when diagnosed early (i.e., cancer that is limited to the submucosal layer) it is curable with an endoscopic mucosal dissection or a minimally invasive surgery. The difference between these two outcomes is time. The earlier the cancer is detected, the better the prognosis.
Currently, upper endoscopy is the primary screening technique for detecting precancerous lesions as well as gastric cancer in the early stages. This technique has a number of downsides: it is invasive, it can have serious side effects (although these are uncommon), and it is expensive and highly dependent on the skill of the endoscopist. For these reasons, endoscopy screening is likely to suffer from poor participation rates. In addition, endoscopy is not a practical approach in low-income countries. There is clearly a need for a less invasive, sensitive screening test that will detect gastric cancer at an early stage.
In an article published in the World Journal of Gastroenterology, Kalnina, et al. reviewed the current status of blood-based biomarkers that: 1) can be used to screen for gastric cancer and 2) can be used to monitor for residual or recurrent cancer. The criteria for markers for these two uses are similar but also somewhat different. According to the authors:
A screening biomarker will:
“be stable and robustly measurable in plasma or serum using routine laboratory equipment, appear in the blood stream before the clinical signs and symptoms arise, should discriminate between cancer and inflammatory diseases and should have a high positive and negative predictive value.”
In contrast, the biomarker used to detecting residual or recurrent cancer:
“must reflect the tumor dynamics. For example it should be rapidly cleared from circulation after complete tumor removal, and it should be able to detect incompletely-resected tumors and to increase in the circulation before the clinical signs of recurrence.”
Analysis of the complex collection of proteins found in blood samples offer a great deal of information including possible biomarkers for cancer. Proteomic analysis involves protein extraction and separation using one of several techniques such as 2-deminsional (2D) gel electrophoresis, enhanced laser desorption/ionization (SELDI) and liquid chip. Protein separation is followed by protein identification by Mass Spectrometry (MS) and bioinformatics. Next the identified proteins are verified using a conventional technique such as Western Blot or ELISA.
There have been a number of promising serum proteomic biomarker studies. For example, in 2004, Ebert et al. (3) used SELDI-TOF-MS coupled with a protein chip technology and a pattern matching algorithm to create a classification technique that achieved 100% sensitivity and 96.7% specificity. This method could also detect early stage gastric cancer with a sensitivity of 89.9%. Liu, et al. (4) used HPLC and LC-MS/MS and three biomarkers (Fibringoen a-chain, apolipoprotein A-II and apoliopoprotein C-I) to distinguish cancer patients from the control group with 93.85% sensitivity and 94.34% specificity. Additional studies using SELDI-MS applications that have shown high sensitivity and specificity ( >90% and >80%, respectively). However, these results still need to be validated using larger studies and the SELDI-MS results have low reproducibility and are inconsistent between research groups.
Although studies have identified promising proteomic, most of the proteins identified are highly abundant plasma proteins that are often part of the coagulation or inflammation process, and many are associated with other cancer types. There needs to be large-scale validation studies to evaluate these biomarkers. In addition, as technical advances improve protein detection sensitivity, new markers could be discovered.
Autoantibodies
Antibodies are interesting as biomarkers because the immune system senses cancer long before the disease manifests itself in other ways. Although studies have shown that autoantibodies against specific tumor associated antigens (TAAs) can be found in the blood up to five years before a clinical diagnosis, there is growing evidence that any individual cancer-associated autoantibody biomarker would have limited diagnostic use. The autoantibody population in cancer patients are diverse and the frequency of individual antibodies can vary widely.
In the studies evaluated by Kalnina et al., autoantibodies distinguished gastric cancer patients from the control population with a high degree of specificity (87–100%) but with wildly variable sensitivity (19.3–98.9%). It is difficult to make a determination as to the clinical usefulness of these biomarkers for population screenings because these studies used different methods for antibody detection, different levels of antibody multiplexing and different criteria for normalization and analysis. In addition not all the studies addressed the issue of overlap between the autoantibody populations in gastric cancer patients and those with benign gastric lesions.
While they may not be effective as screening biomarkers for the general population, autoantibodies could play a role in sorting patients within higher risk groups. Unlike other biomarker types, the immune system can not only sense a tumor at a very early stage, but it can also mount an early high-titer immune response, whereas other biomarkers only increase in circulation as the disease progresses.
Circulating Cell-Free DNA
Cell-free nucleic acids (DNA and RNA) were first observed in human blood in 1948. Although these snippets of biological information come from a number of sources including normal cell death events, several studies have identified an elevated concentration of circulating cell-free DNA (ccfDNA) in plasma of gastric cancer patients. However, elevated levels of ccfDNA is also detected in patients with inflammatory disease, infections and cardiovascular disease, as well as in healthy individuals after exercise. To be useful in gastric cancer diagnosis, the ccfDNAs detected must be specific to gastric cancer. To achieve this, researchers look for gastric cancer-specific sequences, such as MYC and HER2, within the population of ccfDNA found in blood samples. Studies using qPCR showed that an increase in HER2 levels or MYC/GAPDH ratios in plasma distinguished between gastric cancer patients and health control samples. Unfortunately, other studies have found that HER2 levels in tumor tissue does not translate to plasma HER2 levels (5). As these results highlight, it is unclear if gene copy numbers in tumor tissue are consistently and accurately reflected by the ccfDNA levels in plasma.
These assays are also limited by their use of a PCR-based approach. For PCR-based assays to be successful, the patients must have the genetic mutation/sequence targeted by the assay. For that reason, these assays show less promise as routine diagnostic tests. However, targeting specific gene sequences can be useful to identify the presence or absence of a therapeutic target and to monitor treatment progress.
Hypermethylated ccfDNA
Cancer-associated hypermethylation of DNA, including ccfDNA, is a growing area of interest in cancer diagnostics. Several promising methylated markers for gastric cancer have been identified. These include RPRM, XAF1 and a combination of KCNA4 and CYP26B1 (2). In these studies, the sensitivity ranged from 83.9–95.3%, with specificity in the 90% range. Aside from the need for large studies focused on these markers, there are technical aspects of these assays that would hinder their use as screening assays. Currently most methylated DNA studies use sodium bisulfate treatment of the DNA, which converts unmethylated cytosine residues into uracil but leaves the methylated cytosine residues unaltered. Following treatment, the DNA is analyzed with methylation-specific PCR. Unfortunately, incomplete conversion of the unmethylated cytosine residues makes these techniques subject to false-positive results. Recently, several new techniques for quantitative methylation detection have been developed, but to date none of these newer assays have been evaluated in terms of their usefulness in a clinical setting.
Cell-Free RNA
Circulating cell-free RNA (ccfRNA), especially miRNAs, appear to be surprisingly resistant to degradation from RNase activity, extreme pH levels and even freeze-thaw cycles. Several studies have shown that serum from patients have unique patterns of disease-specific miRNAs that are absent from the serum of disease-free individuals. Over 20 studies have explored at miRNA detection as a diagnostic tool for gastric cancer. Some of these studies focused on miRNAs previously identified in gastric cancer tissue samples, others used high-throughput techniques such as microarrays and deep sequencing to identify unique circulating miRNAs. The results for many of these studies have been promising. By comparing pre- and post-operative samples, scientists have identify two miRNAs, miR-451 and miR-486, that decreased in 90 and 93% of post-operative patients, respectively (6,7). Interestingly, other studies found that these miRNAs had a lower diagnostic ability for detecting early stage gastric cancer. This demonstrates that different miRNAs might have different uses in the diagnosis and monitoring of gastric cancer. For early disease detection, the most promising miRNA identified are three miRNA markers that show an increasing expression trend beginning around 15 years before gastric cancer diagnosis. This 3-miRNA panel classified serum samples 2–5 years before clinical diagnosis with an accuracy of 79.3% (8).
Unfortunately, studies to date have had little overlap in the targeted miRNAs, and the results of these studies have been difficult to replicate. For these studies to be effective and informative, there is a need for consistent reporting practice including a consensus on housekeeping genes that can be used as internal controls.
Circulating Tumor Cells and Extracellular Vesicles
Detecting tumor cells circulating (CTC) in the peripheral blood of cancer patients could be a promising method for diagnosis and monitoring gastric cancer. This approach is still in its infancy though, and improvements are need in both the sensitivity of detection and the understanding of CTC biology before these marker can be evaluated for gastric cancer screening and diagnosis.
Interest in cancer-derived extracellular vesicles (EVs) as cancer biomarkers is growing. Currently researchers are still investigating their biological function and composition. These vesicles are either shed or secreted from cancer cells and are found in elevated levels in circulation. They show promise as a source for biomarkers because they carry cancer-derived lipids, proteins, mRNAs and RNAs, and even occasionally genomic DNA, all of which at least partially reflects the parental cancer cells. However, there is little data on circulating EVs and gastric cancer, and no attempt has been made to assign any diagnostic value to EVs for gastric cancer.
Conclusions
Gastric cancer, like many other cancers, earns its deadly reputation largely because it is rarely detected in the early stages. When detected and treated early, gastric cancer has a high survival rate. Unfortunately it is almost asymptomatic until the disease has progressed to an advanced stage. Currently, the most common screening methods for gastric cancer are invasive, resulting in a lower participation rate, and expensive, making access in lower income regions difficult.
Clearly there is a need for a noninvasive diagnostic assays that is sensitive and specific. This is why biomarkers are interesting as cancer screening targets. Over the last decade there has been significant progress in the study and identification of blood-based biomarkers for gastric cancer. Some of these markers have been highly specific and sensitive in early studies. These biomarkers offer the tantalizing promise of one day replacing current endoscopy-, X-ray- and biopsy-based approaches for gastric cancer diagnosis.
References
- WHO: Cancer fact sheet No. 297. Accessed November 23, 2015 (http://www.who.int/mediacentre/factsheets/fs297/en/)
- Kalnina, Z. et al. (2015) Emerging blood-based biomarkers for detection of gastric cancer. World J. Gastroenterol. 21, 11636–53.
- Ebert, M.P, et al. (2004) Identification of gastric cancer patients by serum protein profiling. J. Proteome Res. 3, 1261–6.
- Liu, W. et al. (2012) Proteomic identification of serum biomarkers for gastric cancer using multi-dimensional liquid chromatography and 2D differential gel electrophoresis. Chim. Acta. 413, 1098–106.
- Lee et al. (2013) Clinical significance of intratumoral HER2 heterogeneity in gastric cancer. J. Cancer 49, 1448–57.
- Liu, H. et al. (2012) Genome-wide microRNA profiles identifymiR-378 as a serum biomarker for early detection of gastric cancer. Cancer Lett. 316, 196–203.
Kelly Grooms
Latest posts by Kelly Grooms (see all)
- New Study Suggests Cancer Research Has an Age Problem - April 7, 2026
- Polar Bears, Shrinking Sea Ice and a Scientific Surprise - February 5, 2026
- From Forever Chemicals to Ancient Proteins: Five Science Stories from 2025 - January 8, 2026