As the number of children diagnosed with autism spectrum disorder (ASD) continues to rise, the search for a cause continues. Scientists have been studying genetically modified oxytocin receptors, which have shown promise as a target for studying ASD-related behaviors. One of the obstacles to designing robust scientific experiments for investigating potential ASD causes or treatments is the lack of a truly appropriate model organism for social behaviors in humans (1). Sure, there are the traditional lab rats and lab mice that demonstrate a certain level of social behaviors. However, there has been a loss of natural social behaviors in common lab mice strains because of the reduction in genetic complexity from inbreeding and adaptation to captivity (2). These animals cannot fully represent the depth of human social behaviors, including the ability of humans to form lasting social bonds (1).
Enter: The prairie vole (Microtus ochrogaster).
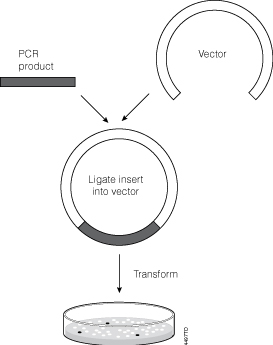
Continue reading “Targeted Gene Modification in Prairie Voles Using CRISPR and pGEM®-T Easy Vectors”