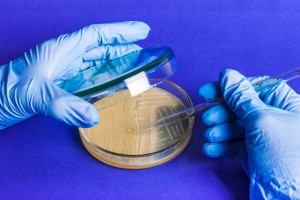
It is estimated that all the bacterial species so far described represent only a tiny fraction of the total. The rest remain unknown to science because they are “unculturable” in standard (or known) laboratory media. Given that many antibiotics were first isolated from environmental bacteria, it seems likely that these as yet unknown organisms could also be a rich source of potential new drug candidates. The desperate need for new strategies to combat multi-drug resistant infections gives impetus to studies investigating how we can culture some of these “unculturable” bacteria and uncover their potential as a source of much-needed new treatments.
Continue reading “Culturing the Unculturable Bacteria”antibacterial
Hope for Treatment of Carbapenem-Resistant Bacteria
But the reservoir of natural products with the potential to act as antibacterial drugs has not yet been exhausted. In contrast to general thinking by drug companies, screening for such products may well still have a bright future” Nature News and Views: “Antibiotic resistance: To the rescue of old drugs” Meziane-Cherif & Courvalin, Nature 510, 477–478.
The emergence of bacteria that are resistant to antibiotics has been an object lesson in the relentlessness of natural selection; the moment a new antibiotic is developed and introduced, the countdown to the emergence of resistance begins. The race to keep the one step ahead of emerging resistance mechanisms has been going on since antibiotics were first introduced.
The history of the development of penicillin and related antibiotics is both an illustration of the ingenuity of scientists and of the never-ending nature of this battle with emerging resistance. The Nature paper is the latest installment in that story. Continue reading “Hope for Treatment of Carbapenem-Resistant Bacteria”
Just a Spoonful of Honey is Medicine Enough
As we face more challenges when treating and healing humans, revisiting therapies that fell out of favor has become more common. For example, people with open wounds that are not healing receive judicious applications of maggots to remove necrotic tissue and promote healing. Leeches are used for patients after surgery to prevent blood clotting in swollen tissues and encourage healing. However, not all therapies involve direct application of squirmy creatures to skin; in fact, honey is one item people are willing to have in their homes. Honey has been used as a sweetener for food, for soothing sore throats and coughs and, more recently, for treating wounds unresponsive to traditional antibiotic therapy. In a recent PLoS ONE paper, the authors assessed the properties that are associated with honey’s antimicrobial activity against pathogenic and food-spoiling bacteria. Continue reading “Just a Spoonful of Honey is Medicine Enough”