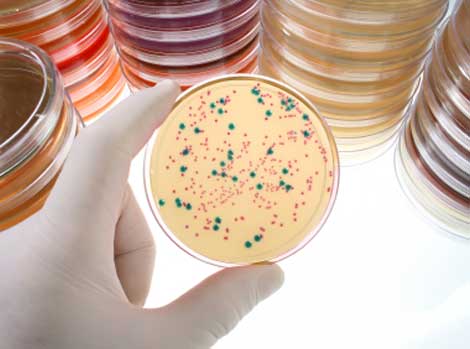
Ah, the wonders and frustrations of cloning. We’ve all been there. After careful planning, you have created the cloned plasmid containing your DNA sequence of interest, transformed it into bacterial cells and carefully spread those cells on a plate to grow. Now you stand at your bench gazing down at your master piece: a plate full of tiny bacterial colonies. Somewhere inside those cells is your DNA sequence, happily replicating with its plasmid host. But wait – logic tells you that not ALL of those colonies can contain your plasmid. There must be hundreds of colonies. Which ones have your plasmid? You begin to panic. Visions of yourself old and grey and still screening colonies flash through your mind. At the next bench, your lab-mate is cheerfully selecting colonies to screen. Although there are hundreds of colonies on her plate as well, some are white and some are blue. She is only picking the white colonies. What does she know that you don’t?
The wonders of lacZΔ E. coli
The process of colony selection can be simplified by choosing a vector and E. coli strain that are compatible with blue/white colony screening. E. coli strains are described as having a lacZΔ when they carry a mutation that deletes part of the β-galactosidase (lacZ) gene. The remaining portion of the gene is called the ω-fragment. By using a plasmid that contains the deleted portion, or α-fragment, the function of the β-galactosidase gene can be restored once the plasmid has been incorporated into the bacterium. For blue/white colony screening, the plasmids have a multiple cloning region within the coding sequence of the α-fragment. When a sequence is inserted into this cloning region, the reading frame is disrupted, and a non-functional α-fragment is produced. This fragment is incapable of α-complementation. Growing the transformed bacteria on a plate containing 5-bromo-4-chloro-3-indoyl-β-D-galactopyranosidase (X-gal) will allow you to distinguish between bacterial colonies formed from cells that contain plasmid with insert from those containing plasmid without insert. Any colony containing the plasmid (and therefore the functioning β-galactosidase gene) will turn blue, a result of the β-galactosidase activity. This is called α-complementation. Those colonies containing plasmids with an insert can be differentiated from those without an insert by the color of the colony (white versus blue). The insert disrupted the β-galactosidase gene, and therefore these colonies remain white. Colonies that did not pick up any plasmid at all will also appear as white colonies; however, most plasmids contain an antibiotic resistance gene that can be used for selection (see below).
There are a number of strains including JM109, DH5α and XL-1 Blue that have the necessary deletions and can be used for blue/white colony screening. However, the mechanism for blue/white screening is slightly different for JM109 and XL-Blue. Both of these strains also have a second mutation, laclq, which increases production of the lacl repressor that stops transcription from the lac operon, and thus production of the α-fragment, until a substrate is present. The substrate, the non-cleavable lactose analog, isopropyl-β-D-thiogalactopyranoside (IPTG), relieves the repression of the lac operon and allows transcription to occur. These strains will need to be grown on medium containing IPTG as well as X-gal.
Occasionally, colonies will appear pale blue, not white. As long as you see colonies on your plate that are darker blue, try picking some of the pale blue colonies, chances are good that they have the constructed plasmid that contains your DNA fragment.
In addition to the β-galactosidase marker, most cloning plasmids will also contain a gene that confers resistance to an antibiotic such as ampicillin. Using Ampicillin (or other appropriate antibiotic) in your growth medium should prevent bacteria that did not take up the plasmid during the transformation from growing. This way you can be fairly confident that the white colonies you see on your screening plate contain plasmid with insert.
And of course it is always a good idea to run controls with your cloning experiment. A plasmid-only control should give you a plate of blue colonies, and this will let you know that your transformation worked. To make sure that the antibiotic in your selective medium is effective, plate some untransformed cells. Few, if any, colonies should be observed on this plate.
Want to learn more and become a true rock star at subcloning? Download the Subcloning Notebook.
Kelly Grooms
Latest posts by Kelly Grooms (see all)
- New Study Suggests Cancer Research Has an Age Problem - April 7, 2026
- Polar Bears, Shrinking Sea Ice and a Scientific Surprise - February 5, 2026
- From Forever Chemicals to Ancient Proteins: Five Science Stories from 2025 - January 8, 2026

I’ve picked some pale colony and made PCR colony. The result is good. It gives the right band of insert. But after 24 hours, the pale blue colony turn into blue colony. I don’t know why. Can you explain it for me.
Thanks so much
Hi,
I seem to remember seeing similar things happening on my cloning plates. I could find a couple of reasons you might see this.
1) The plasmid DNA could be degrading over time, and as a result, all your colonies will slowly turn to blue.
2) Your fragment could be cloned in-frame with the lacZ gene. Such fragments are usually a multiple of 3 base pairs long (including the 3´-A overhangs) and do not contain in-frame stop codons. There have been reports of DNA fragments up to 2kb that have been cloned in-frame and have produced blue colonies. Even if your PCR product is not a multiple of 3 bases long, the amplification process can introduce mutations (deletions or point mutations) that may result in blue colonies.
Pale colonies sometimes result when the insert of interest only partially deactivates the lacZ gene. Since your pale colonies seem to contain the correct insert, I suspect that this is the case for you.
If you have more questions, our technical services scientists are always happy to help!
techserv@promega.com
Hey there! I’m a biochemistry student and I’m taking a Molecular Biology course which is awesome! We performed a gene cloning and transformation on pGEM 9Zf (-) vector with a plasmid, and we inserted an isolated DNA plasmid at the lacZ site. This should have provided antibiotic resistance against ampicillin as well as preventing beta-galactosidase production. What was interesting is that we grow this transformed bacteria on an LB/X-Gal/AMP plate…which presented blue colonies. My inference was that the lacZ gene must not have been completely disrupted, and somehow ampicillin resistance was presented. What would you suggest? It’s getting me confised!
Hi Elie,
Let me see if I have this straight, did you insert a piece of DNA into the multiple cloning region of the pGEM 9Zf- plasmid? If so one of two outcomes can be expected: some plasmid will not take up the insert and will retain a complete lac Z gene; some plasmid will take up the insert, disrupting the lac Z gene.
All of the pGEM 9Zf- plasmids will contain the intact ampicillin resistance gene and will confer ampicillin resistance to any bacteria harboring them. The reason you grow your transformed bacteria on amp-containing medium is to select for bacteria that took up the plasmid during your transformation. The reason X-Gal is included in the medium to differentiate the bacteria that have plasmid containing your insert (white colonies) from bacteria that do not contain insert (blue colonies). You would expect some blue colonies and some white colonies from any cloning, ligation/transformation experiment. If you got only blue colonies, it is possible that something went wrong in the cloning or ligation reaction. What did your positive and negative control plates look like?
Michele
Hi
for me i don’t have bacteria in the plate these the third time, neither white nor blue that mean the plasmid is not inside the bacteria.
I change the protocol three time
Hello, We highly recommend that you contact Promega technical services so that you can speak directly with a scientist who can help you troubleshoot your experiment. They are available at https://www.promega.com/support/customer-and-technical-support/
It is great blog post. I am Always read your blog. Helpful and Informative blog. Thanks for sharing these information with us.
I am a volunteer doing river testing. We are trying to track E-Coli levels. My problem is that differentiating an E-Coli colony from a fecal bacteria colony is not as cut and dry as I would hope. I believe that if the colony has any blue associated with it, it is E-Coli. My group does not offer me enough guidance. https://1drv.ms/u/s!Aidr3l8zer_moGrWKv2DXvcLeZZk Is a link to an image on my one drive account of one of the incubated slides. As an example there is a colony at 5 o’clock close to the outside of the circle that appears to have a very slight hint of blue. Compare this to the colony at 8 o’clock close to the outside of the circle which has no blue at all. Any guidance would be appreciated. Thanks … Michael Bachrach
Hi Michael,
This blue-white screening process refers to subcloning experiments in the laboratory in which you are looking for bacterial cells that have taken up a specific plasmid with a marker. To differentiate bacteria from water samples would be a different process. Our technical services scientists might be able to give you more suggestions, you can reach them at: https://www.promega.com/support/tech-support/
Michele
Assume you used a plasmid that contains the lacz gene in the restriction enzyme site. The plasmid has an antibiotic resistance gene. Following transformation, you grow up the cells on an agar plate containing the antibiotic. What is the result ???
This blog describes how this kind of selection is performed and is not a description with results of an actual cloning experiment.
Yes it uses lacZ for the blue/white selection, and as stated in the blog “In addition to the β-galactosidase marker, most cloning plasmids will also contain a gene that confers resistance to an antibiotic such as ampicillin. Using Ampicillin (or other appropriate antibiotic) in your growth medium should prevent bacteria that did not take up the plasmid during the transformation from growing.”
The antibiotic you will need for selection will depend on the specific plasmid vector you are using in your experiment.