This is the third post in a series of blogs on quantitation for NGS applications written by guest blogger Adam Blatter, Product Specialist in Integrated Solutions at Promega.
Fluorescent dye-based quantitation uses specially designed DNA binding compounds that intercalate only with double stranded DNA molecules. When excited by a specific wavelength of light, only dye in the DNA-bound state will fluoresce. These aspects of the technique contribute to low background signal, and therefore the ability to accurately and specifically detect very low quantities of DNA in solution, even the nanogram quantities used in NGS applications.
For commercial NGS systems, such as the Nextera Rapid Capture Enrichment Protocol by Illumina, this specificity and sensitivity of quantitation are critical. The Nextera protocol is optimized for 50ng of total genomic DNA. A higher mass input of genomic DNA can result in incomplete tagmentation, and larger insert sizes, which can adversely affect enrichment. A lower mass input of genomic DNA or low-quality DNA can generate smaller than expected inserts, which can be lost during subsequent cleanup steps, giving lower diversity of inserts.
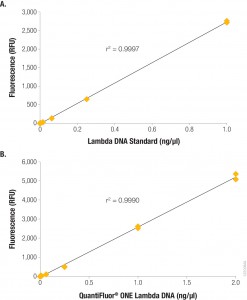
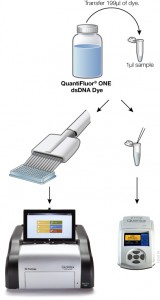
Fluorescent dye-based quantitition is only slightly more complex than a simple absorbance measurement. A standard curve is generated with known concentrations of DNA, and the concentration of DNA in the experimental sample is interpolated from the curve. If using a single-tube fluorometer, such as the Quantus™ Fluorometer, only a blank and one standard are used to calibrate the instrument. For fluorescent quantitation, this measurement is possible over a dynamic range for those concentrations that are common for NGS applications (0.01–400 ng/µl). Much progress has been made in the usability of fluorometers and dye workflows, some of which are available in a truly “add-and-read” format. In these systems, dye solution and as little as 1µl of sample are mixed and measured on a fluorometer, or for higher throughput the protocol can be scaled for a microplate format.
Fluorescent quantitation is also a quick and effective way to quantitate libraries for multiplexing or just before sequencing. These steps are important since mixing libraries together in disproportionate amounts can results in incosnistent coverage and loss of low-frequency mutation detection. Additionally, inaccurate loading of library on your sequencing platform can result in lower than expected coverage or failed runs. Using an appropriate post-library prep cleanup method is critical for removing primers, unligated adaptors or other contaminants that can leaded to overestimation of final library concentrations is critical. If you are concerned about such overestimation, a more exacting quantitation technique, like quantitative PCR, should be used to quantitate libraries before sequencing.
Read Part 1: When Every Step Counts: Quantitation for NGS
Read Part 2: Nucleic Acid Quantitation by UV Absorbance: Not for NGS
Read Part 4: Real-Time (Quantitative) qPCR for Quantitating Library Prep before NGS
Latest posts by Promega (see all)
- ISO 14001 in Biotech: What It Means for Life Science Researchers - April 2, 2026
- When Cancer Research Depends on Quality RNA: Maxwell® RSC in the Lab - March 30, 2026
- Accelerating Drug Discovery at Grove Biopharma with MyGlo® and ProNect® - December 9, 2025

In the article you said “Fluorescent dye-based quantitation uses specially designed DNA binding compounds that intercalate only with double stranded DNA molecules. When excited by a specific wavelength of light, only dye in the DNA-bound state will fluoresce.”
So only dsDNA can be quantified with DNA-dye based fluorescence system? I thought you can measure dsDNA as well as ssDNA using fluorescence method
It depends upon which DNA binding dye you use. We sell separate dyes for ssDNA and dsDNA https://www.promega.com/products/dna-purification-quantitation/dna-and-rna-quantitation/