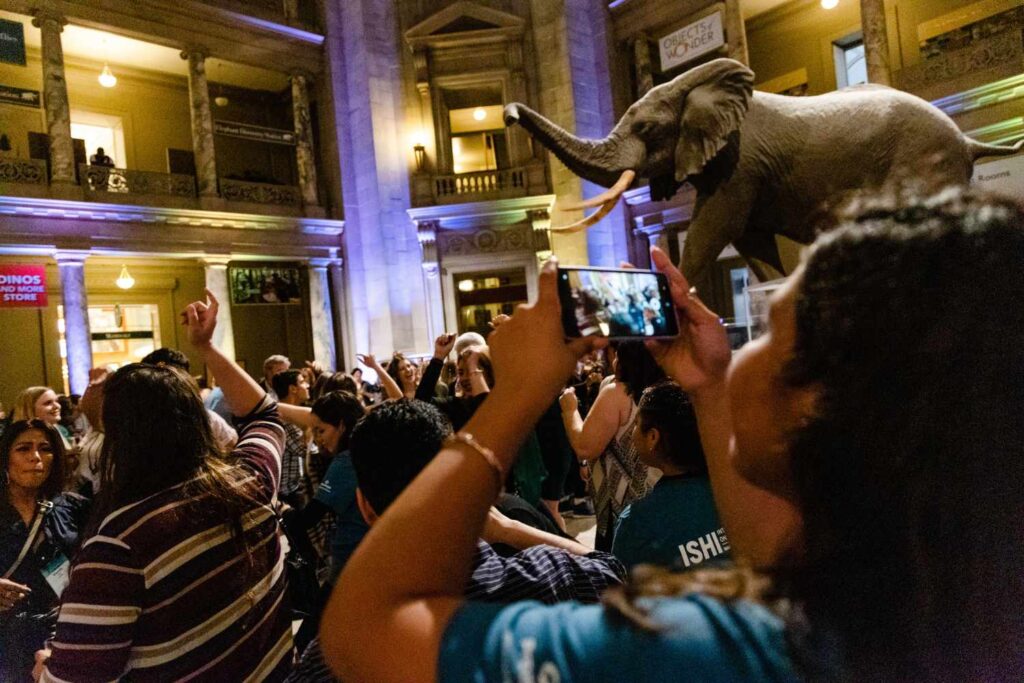
It’s hard to imagine a better way to celebrate the 33rd International Symposium of Human Identification than a night spent wandering through the Hall of Human Evolution at the Smithsonian Museum of Natural History. The meeting, which took place in Washington D.C. from October 31–November 4, focused largely on using investigative genetic genealogy (IGG). When used to identify human remains or solve cold cases, IGG (a.k.a. forensic genetic genealogy or forensic investigative genetic genealogy, take your pick) relies heavily on techniques developed to sequence DNA from ancient human remains.

New to ISHI this year were live-streamed presentations, building off the success of last year’s session recordings for online streaming. Another first was attendees dressing up in costume for the welcome reception, which happened to coincide with Halloween. From a nucleic acid-themed group costume to Sims characters to a bunch of grapes, ISHI 33 attendees had a chance to show off their fun side while reconnecting with colleagues.
While a range of topics were covered during the workshops, sessions and poster presentations, three themes stood out to this first-time ISHI attendee. In addition to IGG, there was widespread interest in developments in DNA databases as well as efforts to mobilize DNA analysis labs.
Continue reading “The 33rd International Symposium on Human Identification: The Past, Present and Future of Investigative Genetic Genealogy”


 While the forensic and general communities continue to argue about the merits of the recent Supreme Court ruling on collection of samples from arrestees prior to conviction, I am fascinated by the technology that make this question relevant. The conventional way of generating a DNA profile from a sample by STR (short tandem repeat) analysis is a long process involving a series of steps that require sophisticated expensive equipment, trained personnel and, more importantly, time. The actual process of DNA analysis consists of a) sample collection, b) DNA extraction, c) PCR amplification using 16 or more unique fluorescently labeled primer sets d) capillary electrophoresis to size labeled DNA amplicons, e) software analysis to size DNA fragments and allele calls based on migration of allelic ladder fragments, and f) comparison to known profiles in the database. This entire workflow can typically take days or even weeks, and therefore it is not surprising that we see newspaper reports of backlogs of criminal and other property cases. With these time ranges, the sample collected at a site would be of no practical use to most ongoing investigations.
While the forensic and general communities continue to argue about the merits of the recent Supreme Court ruling on collection of samples from arrestees prior to conviction, I am fascinated by the technology that make this question relevant. The conventional way of generating a DNA profile from a sample by STR (short tandem repeat) analysis is a long process involving a series of steps that require sophisticated expensive equipment, trained personnel and, more importantly, time. The actual process of DNA analysis consists of a) sample collection, b) DNA extraction, c) PCR amplification using 16 or more unique fluorescently labeled primer sets d) capillary electrophoresis to size labeled DNA amplicons, e) software analysis to size DNA fragments and allele calls based on migration of allelic ladder fragments, and f) comparison to known profiles in the database. This entire workflow can typically take days or even weeks, and therefore it is not surprising that we see newspaper reports of backlogs of criminal and other property cases. With these time ranges, the sample collected at a site would be of no practical use to most ongoing investigations.